From hibernation to neurodegeneration: how a cold-resistance protein is paving the way to dementia treatments
Hibernating animals endure achingly cold temperatures and emerge unscathed, a phenomenon in part attributable to the protein RBM3. Research in mice has shown that RBM3 expression is protective against misfolded protein damage, suggesting the protein as an interesting target in the development of therapies for neurodegenerative conditions such as Alzheimer’s disease.
How can we mimic the protective effect of RBM3 without needing to cool patients repeatedly and for extended periods of time? To find out, we need to understand how this system is regulated, and what exactly RBM3 does in the cells. In a recently published project, Julie Qiaojin Lin and Deepak Khuperkar found and connected the edge pieces of this puzzle.
Julie is a Sir Henry Wellcome Fellow and Emerging Leader at the UKDRI, and a postdoctoral researcher in Giovanna Mallucci’s research group. Deepak is a joint postdoctoral researcher with Marc-David Ruepp at King’s College London and Giovanna Mallucci at the Cambridge UKDRI.
Where did the idea for this project come from?
Julie. It all started with the Open Targets project led by Giovanna Mallucci and Manos Metzakopian. They wanted to find out how RBM3 is regulated by temperature — these kinds of proteins are known as ‘cold-shock’ proteins. We currently know very little about RBM3, so our screen was a shot in the dark; we had no idea what to expect from it.

What did you find?
Julie.We performed the screen using a CRISPR-Cas9 genome-wide library and knocked out each gene in the library, one gene per cell. We then selected the cells with the most increased or most decreased RBM3 expression, which gave us a set of genes to investigate. Curiously, these followed a theme in molecular function: the regulation of splicing*.
These results were corroborated by an RNA sequencing analysis we carried out in parallel, which revealed that depending on the temperature, RBM3 had different isoforms created by differential self-splicing. RBM3 is an RNA-binding protein, and we needed an RNA biologist, which is when Deepak joined the project.
Deepak. When I joined the lab, the main question was: what does RBM3 do? I was fascinated when I heard about its neuroprotective functions, and was excited to dive into the mechanism.
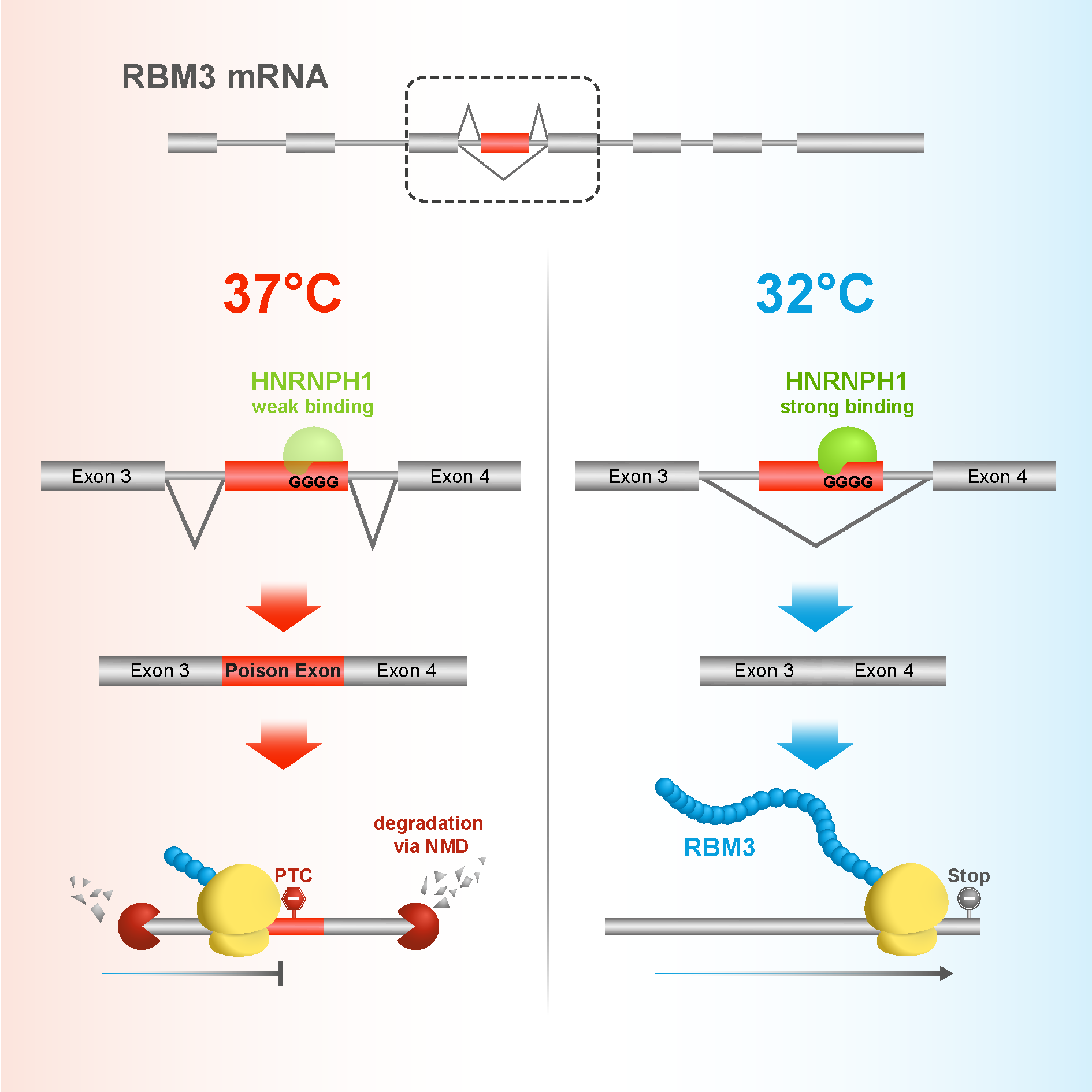
Julie and I realised that there might be something happening with alternative splicing of RBM3. We found that by default, RBM3 contains a poison exon. This means that transcripts of the gene are destroyed before they can be translated into protein. However, when the cells are cooled, the poison exon is spliced out of the transcript, which can then be translated, and levels of RBM3 protein rise in the cell.
To test this further, we looked into the top hit from our screen, a gene called HNRNPH1. When we mutated HNRNPH1 sites on the poison exon, RBM3 was no longer able to respond to cold shock. This brings the story neatly together, showing that in response to cold induction, HNRNPH1 binds to the poison exon on RBM3, which is then spliced out of the transcript, allowing RBM3 proteins to exert their neuroprotective effect. It was a nice mechanistic connection to the initial finding.
*Splicing: the inclusion or exclusion of different parts of the RNA sequence before its translation into protein molecules.

Was there anything surprising about your results?
Julie. We actually didn’t expect anything from the screen. Our best guess was that the regulation was that low temperatures decreased the fluidity of cell membranes that they would be less able to stretch. And so we thought we might find genes related to mechanosensitivity in the screen, for example ion channels. So the fact that our hits were splicing factors was a complete surprise.
Deepak. It was definitely a pleasant surprise — no one knew about this poison exon. And it’s a satisfyingly neat and tidy method of regulation; such a tiny step that does wonders.
Poison exons are part of a larger group of hidden or cryptic exons, so-called because these sequences are hidden among the gene’s introns, the sections of DNA that don’t encode a protein sequence but that are important in the regulation of the gene’s expression.
Julie. RBM3 is quite special though, because in this case the poison exon is regulated by temperature. Most cryptic exons are regulated through classic feedback mechanisms, but RBM3 might be one of the few proteins where temperature affects the poison exon, which in turn affects the gene’s expression.
Deepak. It’s also a rapid response method of regulation. It might seem like a waste for a cell to transcribe a gene only for that transcript to be destroyed by a poison exon, but it’s quite clever — it means that the cell can respond very quickly when there is a change in the environment. So as soon as the cell senses that the temperature is dropping, RBM3 can be active to protect the cell.
Was there anything particularly challenging in this project?
Julie. It was a lot of small experiments, but to be honest most things actually worked surprisingly well!
Deepak. The mini-gene experiment was a tricky one. To test our hypothesis about how RBM3 is regulated, we created an artificial system in the lab. A mini-gene is a piece of DNA sequence which we engineer so that we can easily include or exclude pieces of the sequence and see how that affects the function of the gene. We can express this mini-gene in our test cells, and this is a very quick way to play around with the sequence.
What are the next steps in this research?
Deepak. From this work, we know how RBM3 levels are first increased when the cells are exposed to the cold. The main question now is to understand what happens next, the mechanism through which RBM3 actually protects cells.
Julie. This work has set us up with a lot of tools that we can now use to manipulate RBM3 and dig into this question.
Deepak. And once we have a complete picture, our hope is that this work can be used to develop therapies for neurodegenerative diseases.