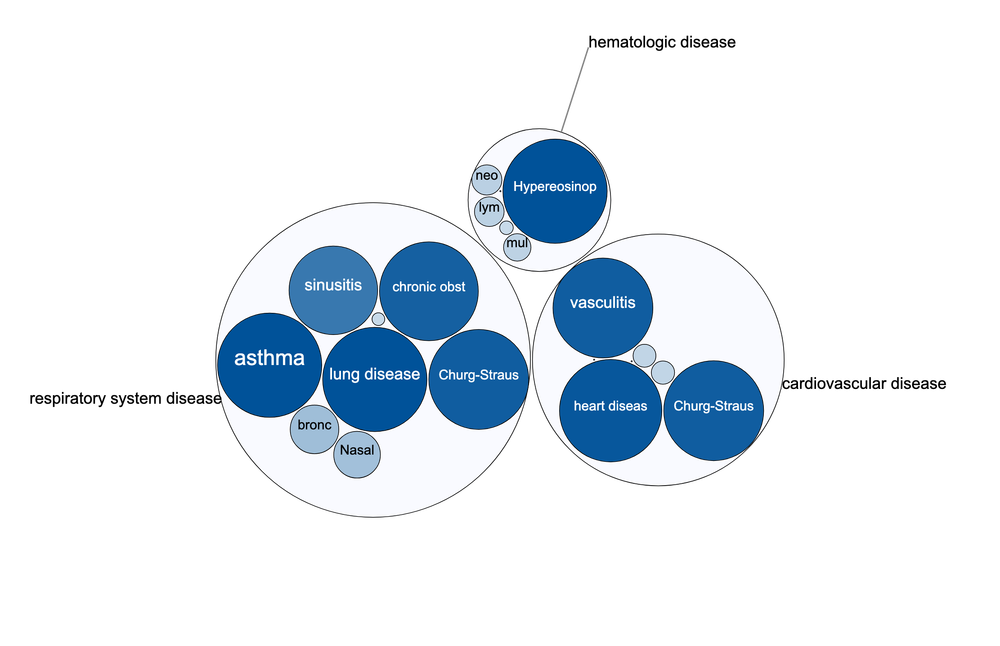
Open Targets Platform: release 19.06 is out
We've just released the latest update of the Open Targets Platform, release 19.06.
What's new?
- Target safety information
- TEPs and chemical probes
- Target-disease associations
Head to release notes or use the REST-API stats endpoint for a breakdown of the numbers of drug targets, diseases, evidence, and target-disease associations per data type and data source.
Target safety information
As a follow-up to the safety data in Open Targets Platform release 19.04, we now have more targets with known safety effects and safety risk information, including TBXA2R and JAK2.
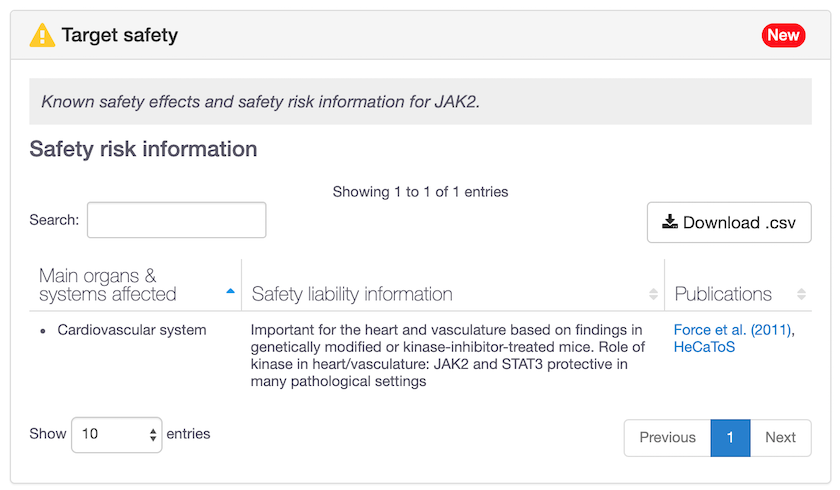
"Is my target safe to be modulated?" Look for known safety effects and safety risk information in the Open Targets Platform, now available for 193 targets.
The known safety effects can vary from drowsiness to increased heart rate, coma or even death, whereas safety risk information mainly includes cardiotoxicity and hepatotoxicity.
Check the Target profile documentation page for the list of papers and website we've curated target safety information from.
TEPs and chemical probes
In this release, we've included the latest Target Enabling Packages (TEPs) for GALT, GALK1 and MLLT1. We've also added more chemical probes, small-molecule modulators of a protein’s function that can be used in cell-based or animal studies.
These are two examples of new chemical probes that are worth noting as they result from a collaboration between SGC and Takeda, one of the Open Targets partners:
All TEPs come from the Structural Genomics Consortium (SGC), whereas chemical probes are from the SGC, the Chemical Probes Portal and/or Open Sciences probes.
If you are looking into probing your next target using chemical probes, you will be pleased to hear about these new additions.
Target-disease associations
A new release always means new evidence available for novel target-disease associations. We've hand picked the following examples for you:
-
Text mining evidence for IL5 in Bronchiectasis
-
Animal model evidence for ADORA3 in glaucoma
-
Somatic mutation evidence for TP53 in Gallbladder Adenosquamous Carcinoma
-
Rare missense variant (rs143718918) from UniProt as evidence for ABCA7 in Alzheimer's disease
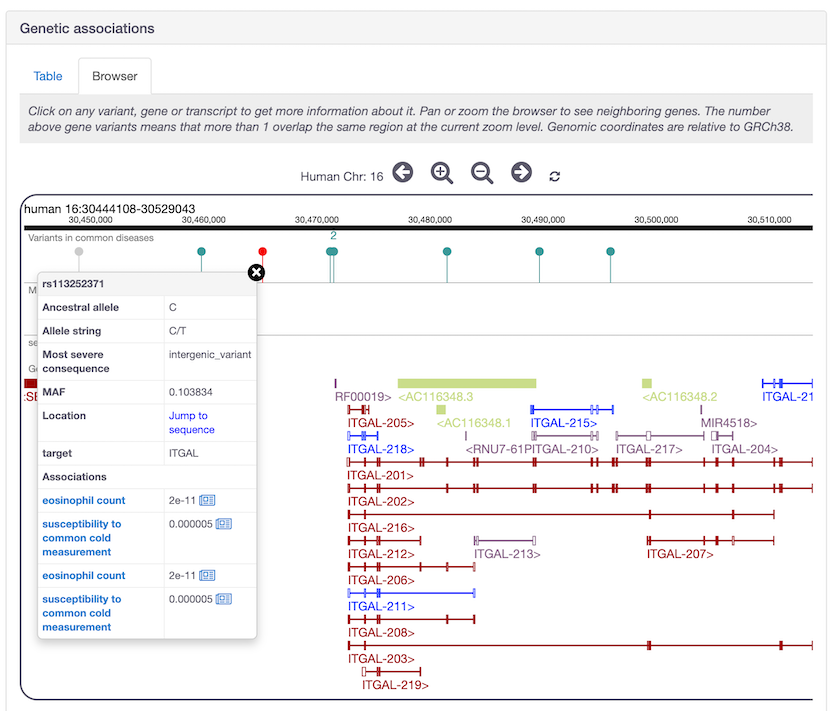
SNP rs113252371 (see the red lolipop above) is one of the new GWAS Catalog common variants now availabe as evidence for ITGAL in eosinophil count and other measurements.
Other news
More entries displayed in the drug table
Did you know that some diseases may have tens of thousands of drugs in clinical trials?
Although the Open Targets REST API is the best way to access this (big) drug data, we've now increased the number of rows displayed on the user interface, so that you have instant access to all drug entries while still navigating an easy-to-use website.
Want to get drug information using the Open Targets REST API but don't know where to start? Email us and we'll help.
If a disease has up to 10,000 drug entries, e.g. asthma with 2,587 entries, the new table will display them all, so that you can explore the data straight away.
But if the number of entries happens to be higher than that, we now cap the row number to 10,000 and provide a download option for you, located at the top of the table.
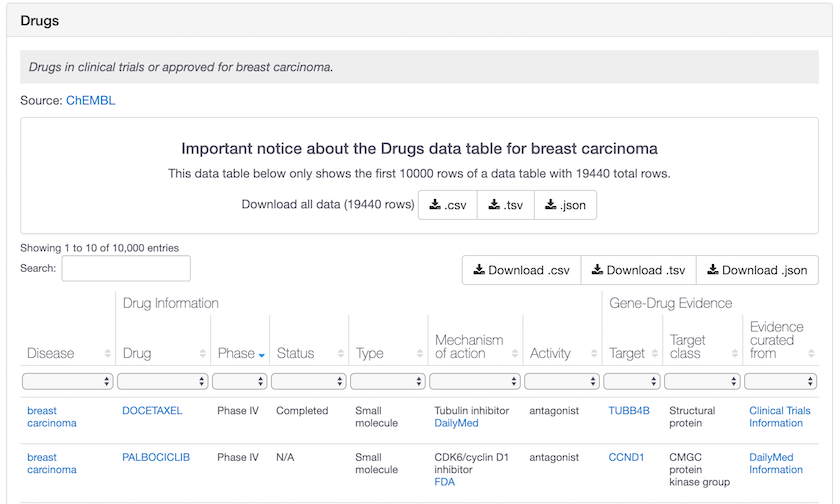
Breast carcinoma, cardiovascular and many other diseases have more than 10,000 drug entries. You can now download this entire dataset in csv, tsv and json. All with a simple click of the mouse!
Please be aware that it may take a little while for the table to load and for the download to complete.
Expression evidence
We've also made changes to the differential expression evidence from Expression Atlas, resulting in much less evidence in release 19.06 when compared to previous releases. We now only:
-
Select the microarray probe that has the highest mean expression per gene
-
Include genes that are differentially expressed in
disease versus normalcomparisons and filter out any other comparisons
Save the date
The next release of the Open Targets Platform, release 19.09, will be three months from now.
Till then, if you have any comments or questions, please email us. You can also join the Open Targets community on Facebook, LinkedIn and Twitter and keep up-to-date with upcoming Open Targets data, publications and job openings.