Our latest release — 16.11 — is out
The latest release of our Open Targets Platform is out.
We have plenty of new web features and new data to help your journey into drug discovery.
These are our latest stats:
- 31,071 targets
- 8,659 diseases
- 2,559,080 target-disease associations
- 4,973,211 evidence from 13 data sources
Want to give input into the future development of our Platform? Please take a moment to complete our survey.
Revamped super duper homepage
Our homepage has now a sleeker look-and-feel and is seamless to navigate through. It is also mobile-friendly and includes a social media hub.
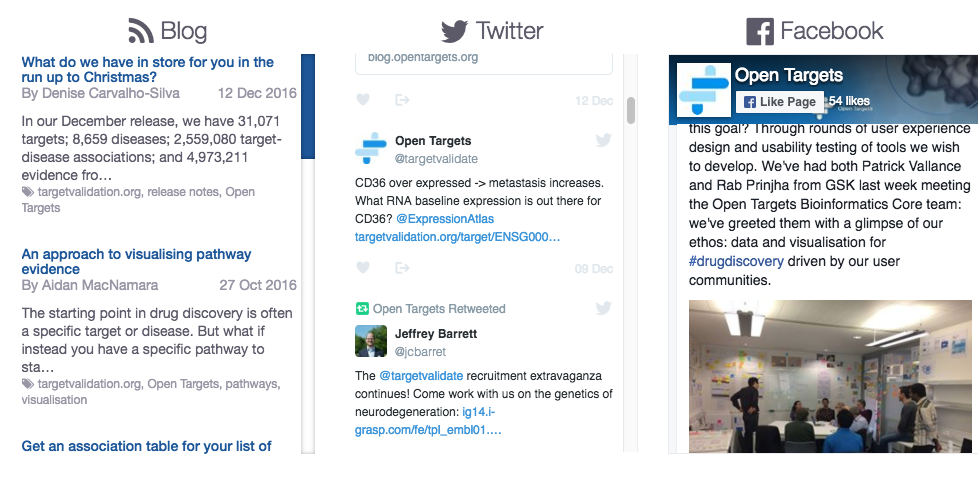
Try our new homepage and find out how to access our data for free, get timely help from us, read our Open Targets story.
Python client
You may have seen our API tutorial and some examples of how to access our data.
We now also offer a Python client, a powerful alternative to access our REST API. If you know Python, give our client a go and let us know how you get on.
Matching targets for drugs or phenotypes
We now match targets and diseases to two new entry points, drugs and phenotypes.
Why not search for humira or hyperlordosis to find out which targets and diseases they match to?
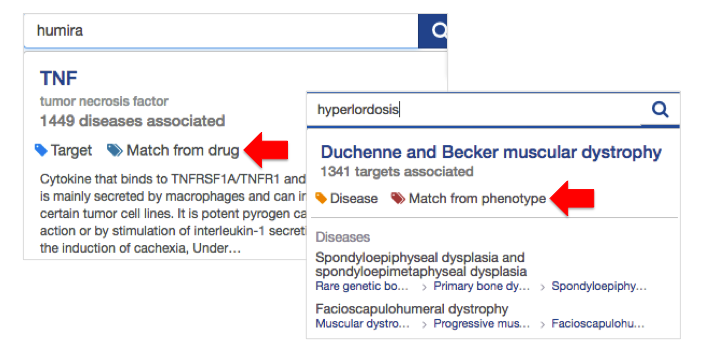
Note the tags "match from drug" or "match from phenotype" in the results of your search.
Your target list
Do you want to restrict the target-disease associations in our Platform to a defined target list?
You can now upload a list of target names (e.g. IL23R, ITGA4) or Ensembl IDs (e.g. ENSG00000162594, ENSG00000115232) for a disease of interest (e.g. inflammatory bowel disease) and get the association scores and the evidence for the association.
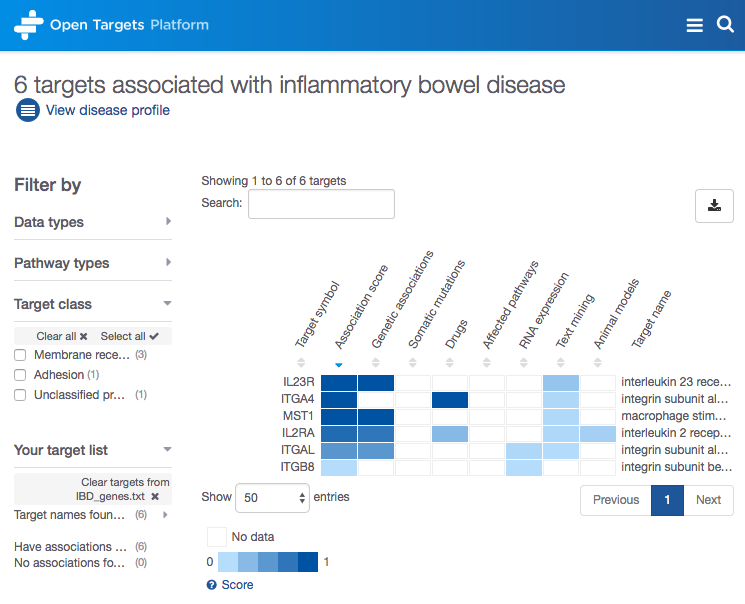
You can clear the selection of your target list at any time you wish.
Filtering targets associated with a disease
Would you like to filter your targets based on whether the protein is an Enzyme (e.g. Kinase, Protease), Membrane receptor (e.g. Toll like) or Structural protein, for example?
Look out for our new web feature 'Target class', make your choice and focus on the types that matter for you.
GTEx visualisation
By popular request, we now display the latest version of the GTEx box plot for baseline expression variability of mRNA.
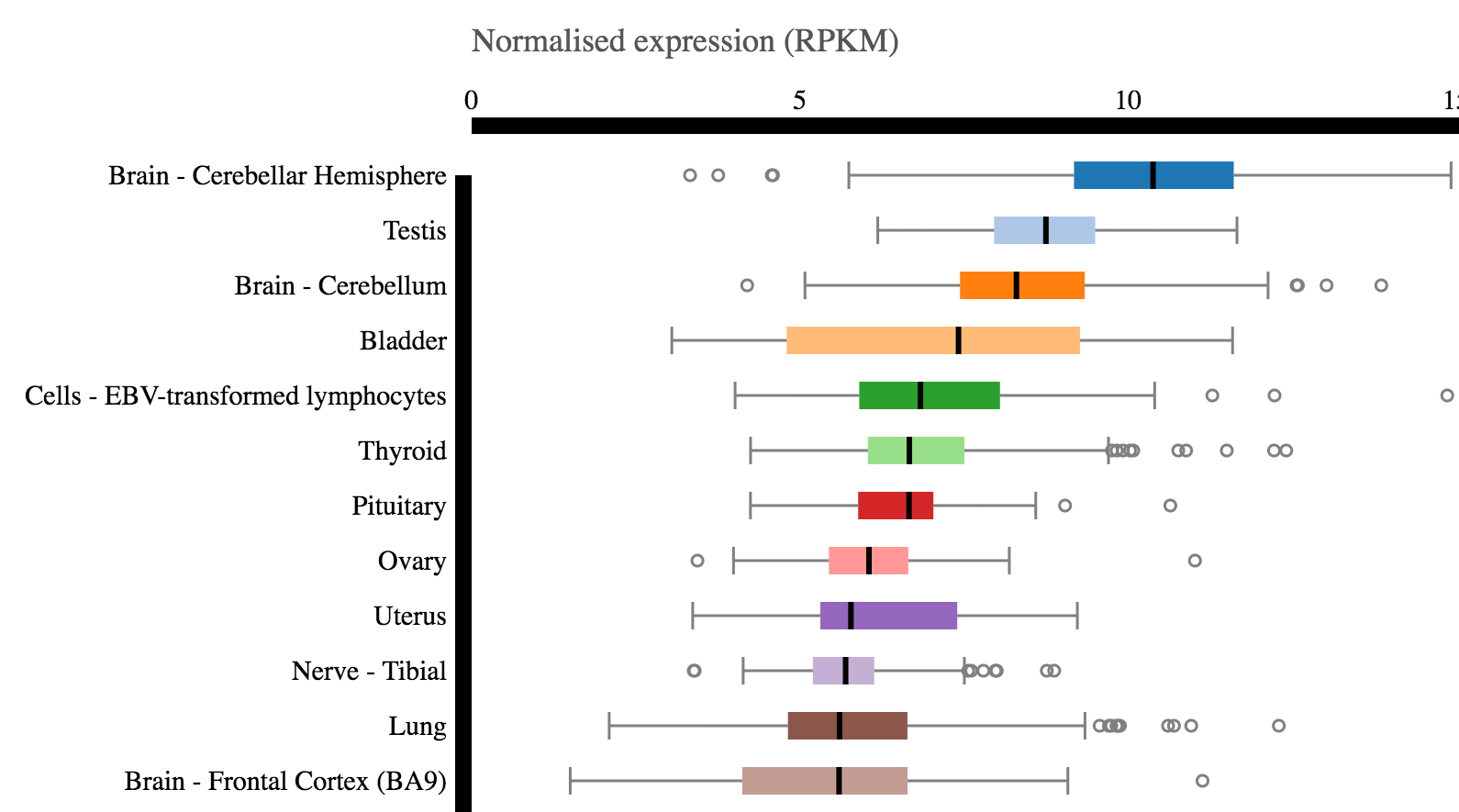
You can still get all other experiments for mRNA baseline expression from Expression Atlas, which also includes GTEx and FANTOM5 besides many other projects.
New infrastructure
We have moved to kubernetes for automating the deployment, scaling, and operations of our Open Targets applications on a Google Cloud Platform. This allows us to roll out new services more easily and it gives flexibility in our agile environment.
Outreach page
Since the launch of our Outreach in Open Targets, we have been providing training workshops and tutorials.
Why not check our Outreach and tutorials page to find out where we will be teaching next? You can also arrange a workshop in your lab (available for both academia and industry).
Want to know more about the Open Targets Platform? These are some of the pages that are a good starting point:
And get in touch with us if you have suggestions or comments on this release or on our Platform.