Open Targets releases our COVID-19 Target Prioritisation Tool
After several months of hard work, we are excited to introduce the Open Targets COVID-19 Target Prioritisation Tool, an interactive, open source web portal that allows you to systematically explore host and viral targets for COVID-19. The tool is available at http://covid19.opentargets.org/.
With the global coronavirus pandemic showing no signs of slowing down, scientists are in a race to develop therapies that can treat and prevent COVID-19. While vaccines are progressing through clinical trial phases, there is also a concerted effort to explore how experimental or approved drugs could be used as treatments (e.g. antivirals or immune-suppressors).
At the start of the pandemic lockdown, our Director Ian Dunham wrote about what we were doing to support research into potential COVID-19 drug targets. Because we focus on systematic target identification and prioritisation, we know that identifying potential drug targets is a complex problem and that clinical trials experience high attrition rates due to lack of drug efficacy or safety. The challenge for COVID-19 is similar and we expect that a high percentage of the ongoing COVID-19 clinical trials will fail for these reasons.
Building on our initial work, we believed that we were in a position to develop a useful tool to aid drug target identification and drug repurposing for COVID-19. To do this, we integrated molecular and clinical data from EMBL-EBI and other public resources to provide an evidence-based framework to support decision-making on potential drug targets and treatments for COVID-19.
Systematic exploration of potential COVID-19 drug targets
To date, Open Targets has focussed exclusively on human targets. To support research into therapeutic approaches to treating COVID-19, we would have to integrate data across both host and pathogen targets to foster a better understanding of host-pathogen interactions against a back-drop of fast emerging results.
As such, we created the Open Targets COVID-19 Target Prioritisation Tool and integrated a number of key public datasets and publications to facilitate systematic exploration of host and viral targets as well as potential treatments for COVID-19.
With powerful search, filter, and sorting capabilities the tool enables the construction of complex queries and you can find targets based on key criteria including:
- Protein interaction and network data providing evidence of direct or indirect interaction with a SARS-CoV-1, SARS-CoV-2, or other viral protein, which includes relevant publications such as Gordon et al., Nature (source: IntAct)
- Up- or down-regulation of protein abundance levels at specific time points during COVID-19 infection (source: Bojkova et al., Nature)
- Clinical trial information including: the number of drugs where the target is identified in the mechanism of action, the maximum trial phase for any drug that modulates the target, and if there are any trials involving these drugs where the indication is COVID-19 (source: ChEMBL)
- In vitro data for chemical compounds assayed in the context of COVID-19 infection mapped to their corresponding targets using the known drug mechanism of action (source: ChEMBL)
- Small molecule, antibody, and other modality target tractability assessments (source: Open Targets)
- Baseline expression, distribution, tissue, and subcellular location data (source: Human Protein Atlas, Expression Atlas)
- Safety information, including known safety effects and non-clinical experimental toxicity data (source: Open Targets)
- Publications where the target and COVID-19 or its synonyms terms co-occur in the same sentence using text mining (source: Europe PMC)
By utilising a mix of the column-specific filters and sorting, you can answer a variety of research questions that can inform therapeutic hypotheses about novel COVID-19 drug targets and opportunities for drug repurposing.
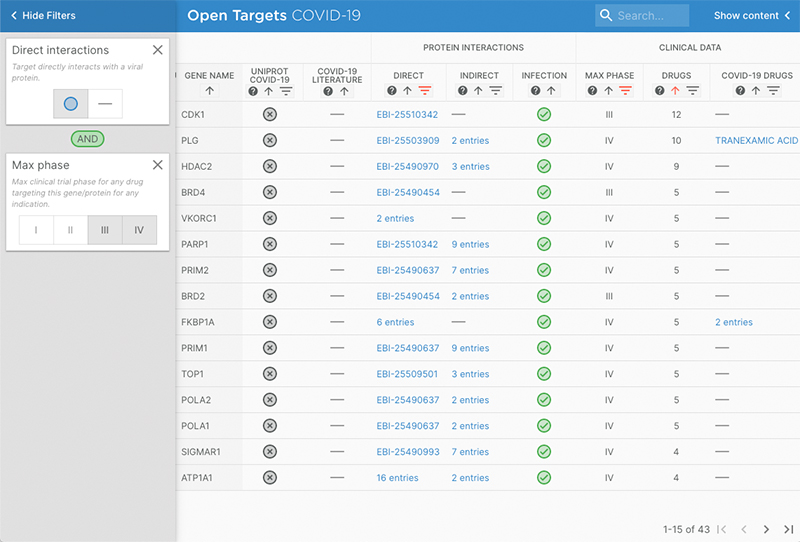
The tool can be used to identify human targets that directly interact with a viral protein and have a Phase 3 or 4 drug
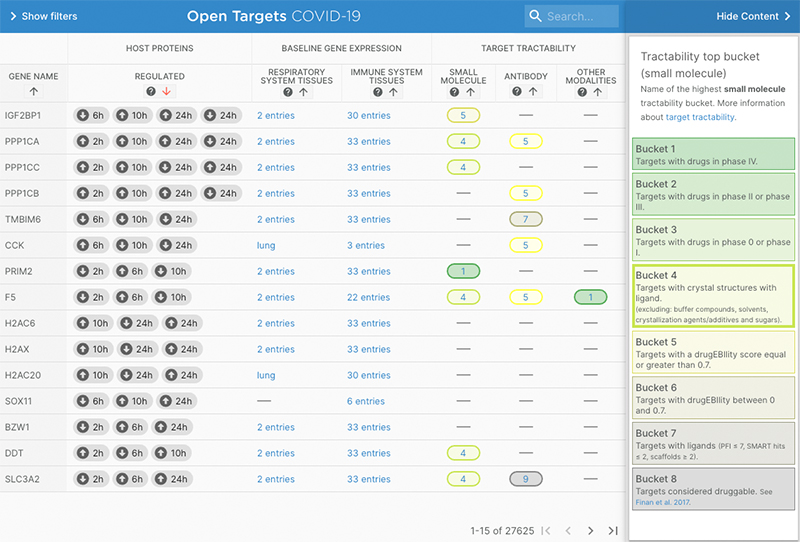
The tool can also be used to discover human targets altered in abundance during SARS-CoV-2 infection and are expressed in the immune system and potentially small molecule tractable
As an alternative to the web interface, you can also download the entire dataset for further analysis. And in keeping with Open Targets philosophy, we have made this project open source and you can clone the data pipeline and web application repositories to integrate and display your own data.
Future plans for the tool
We developed our approach as a small contribution to the research community, hoping our offering is complementary and that users can gain value alongside other approaches (e.g. COVID-19 UniprotKB, Ensembl COVID-19, Coronavirus CanSAR, HPA Chemical Checker). The tool is part of a wider EMBL-EBI response to COVID-19 and has also been included in the COVID-19 Data Portal to facilitate data sharing and analysis and accelerate coronavirus research.
Given the rapid advancements in our understanding of COVID-19 and emerging research, our plan will be to periodically update the portal and integrate new datasets while seeking funding for further development and expanded functionality.
We are interested to hear about other datasets that might help inform the selection of potential COVID-19 targets and how the tool has helped you triage the increasingly abundant information.
If you would like to collaborate on this tool and work with us on expanding it with new datasets and features, please get in touch.




