How Open Targets is investigating 100 chromatin-related disorders | Rare Disease Day 2021
Rare diseases – despite their name – are not uncommon; 1 in 20 people will live with a rare disease at some point in their life. Many of these diseases don’t currently have cures, and as they are little understood, they are often undiagnosed.
Rare disease day is celebrated annually on the last day of February, to raise awareness of rare diseases and how they affect patients’ lives. Several of the experimental projects within the Open Targets consortium investigate rare diseases, including the 100 Chromatin-related disorders project (OTAR2-053).
Sebastian Gerety is a Senior Staff Scientist at the Wellcome Sanger Institute working with Matt Hurles, who co-leads OTAR2-053 with Emmanouil Metzakopian (UK DRI-Cambridge) and Helena Kilpinen (the Wellcome Sanger Institute, UCH London, and the University of Helsinki, Finland). Sebastian oversees the project and, in honour of Rare Disease Day 2021, told us about what they've been working on.
Which diseases are you researching?
Childhood disorders involving traits like intellectual disability, Autism Spectrum Disorder, and delayed milestones affect many thousands of kids worldwide. We use the term Neurodevelopmental disorders (NDDs) because they affect the nervous system and are often caused by problems in the development of the brain. Recent research has highlighted the fact that certain genes affected in NDDs are also affected in psychiatric disorders such as schizophrenia and bipolar disorder.
Projects like Deciphering Developmental Disorders (DDD) at the Sanger Institute (led by the Hurles Faculty group) help us understand that NDDs are often caused by spontaneous mutations around conception ("de novo mutations"). Although projects like the DDD aim to improve diagnosis rates, children with NDDs are still only rarely given a genetic diagnosis in normal clinical settings. Even when a family does find out the genetic cause, this is only one piece of a challenging puzzle for child and parents: little is known about how these genetic changes cause a disorder, and vanishingly few have specific treatments available.
How does this project address the gaps in our understanding of rare diseases?
Our project is based on the premise that by understanding the disorders at a molecular level, we can then seek treatments that fit our new understanding.
In this project we are attempting to recreate the mechanism of numerous NDDs, starting with cells in a dish (Induced Pluripotent Stem Cells — IPSCs) which have been modified to have the genetic changes found in children with the disorders. We can then analyse the cells to gain insights into the effect of disease. We use RNA sequencing of samples to look deeply at the changes occurring in the disease cells.
One nice thing about using cells is that they are often scalable: we can grow cells for many different disorders densely in petri dishes, allowing us to make embryonic brain cells (Neural Precursor Cells — NPCs) from them on a large scale. A single Research Assistant in Sanger's Cellular Generation And Phenotyping facility can usually handle 12 cell lines or more at once. This means that we can simultaneously process cells lines for 5 different disorders, and compare them to the control cell lines.
Because each is individually quite rare, this work can be most impactful by studying many rare disorders at once. In the project, we are examining up to 100 different disorders together, with the exact same methods, looking for similarities between them that give us clues to groups of disorders that might respond to a common therapy.

How does this research lead to new therapies?
The second arm of the project, based in Emmanouil Metzakopian’s lab at the UK Dementia Research Institute, is to actually investigate treatments. Here too, we can use the scale of in vitro cell culture to test as many compounds as we can get a hold of.
We are treating stem cells with drugs and compounds – namely chromatin modifiers – that act on proteins related to those damaged in NDDs. Using bulk RNA sequencing, we can look at the effect of many drugs on embryonic neural cells. We can then try to pair up cells modelling a particular rare disease with a drug that looks like it might counteract the disease effects.
Although we have some interesting drugs and compounds, very few proteins actually have drugs that affect their function. To compensate for this, the Metzakopian lab are performing a larger screen to look at the effect of inactivating 500 different chromatin-modifying genes individually. Using a CRISPR/CAS9 gene editing approach allows us to run the same basic assay repeatedly to work through the effect of 500 different gene inactivations.
Once we identify either a compound or a gene inactivation that counteracts the effect of a disease, we can test our treatment hypothesis by trying it out in our disease cell models. This finding can then be followed up by one of our collaborators in the pharmaceutical industry with the hope of designing a drug that will replicate this effect.
What's in a brain?
Of course, no cell-based model is a perfect representation of a disease mechanism that normally occurs in the complex orchestra of human brain development. To expand our understanding of NDDs, Helena Kilpinen and her group are taking NDD cell lines and models, and turning them into mature neuronal cell types of the brain (cortical neurons).
While these cell types are closer to what we expect in a child's brain after birth, the experiments to generate these brain cell types are challenging to run, take many months to complete, and cannot be handled on the scale of IPSCs and NPCs. Because of this, fewer disorders can be examined using these involved methods. Persistence, effort, and lots of troubleshooting is helping make progress with this difficult part of the project. By borrowing from other Open Targets projects, and as technologies develop, we may find ways of pooling cells from many disorders. This would allow Helena's group to run many more samples at once, and potentially parallel the scale of work done on IPSCs and NPCs.
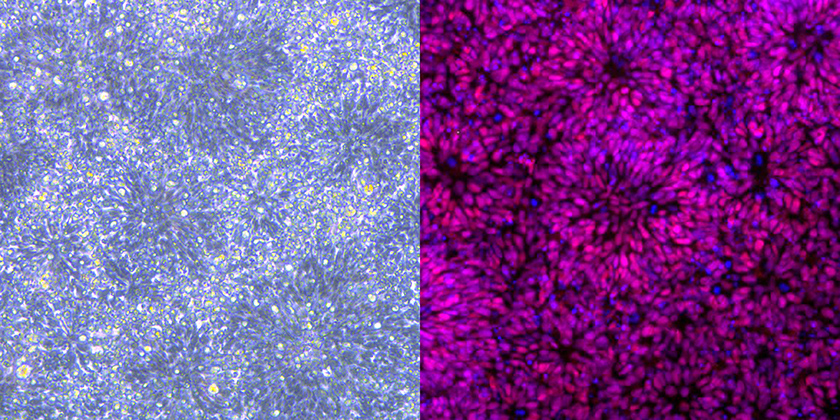
Human neural stem cells that have been generated and stained as part of the 100 chromatin-related disorders project
Where are we now?
While the COVID pandemic has caused a pause in our lab based activities over the summer, we have all managed to restart experiments in September and so the project progresses at COVID pace. Bioinformatics never sleeps, and so has continued throughout lockdown, albeit thirsting for new experimental data to flow from the wet lab.
We are looking forward to further fruitful engagement with the Open Targets family members as the project evolves and opportunities emerge. In particular, as the drug and gene knockout data flows from the Metzakopian group, the opportunity to compare disease and treatment effects will take off, providing us with new avenues of treatment validation to explore.