We've released Open Targets Platform Version 1.2!
That's right. The Open Targets Platform v1.2 is out!
In this release, we have new web displays and plenty of extra data to assist you in drug discovery and validation:
- 30,591 targets
- 9,425 diseases
- 4.8 million evidence
- 2.4 million target-disease associations
New Web Widgets
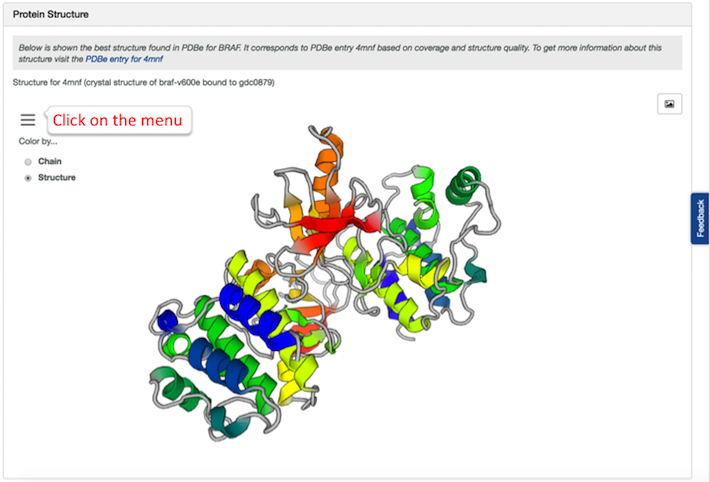
We have up-to-date widgets for both 'RNA baseline expression' and 'Protein Structure' of a target. In the latter, you can now rotate the protein structure, change its colour, zoom in and out, and highlight any amino acid residue:
More Data Sources
We have two additional data sources: somatic mutations from IntOGen and variants curated by clinical geneticists from Gene2Phenotype.
Check our Target Validation Platform Data Sources page for a complete list of sources used in our computational analyses.
Added Model Organisms
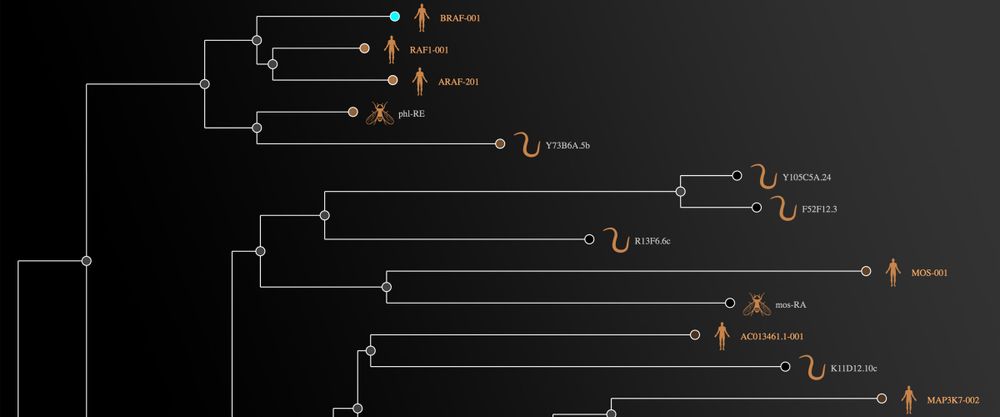
Fly and worm are two of the new species in our phylogenetic trees, which have a new name, 'Gene tree':
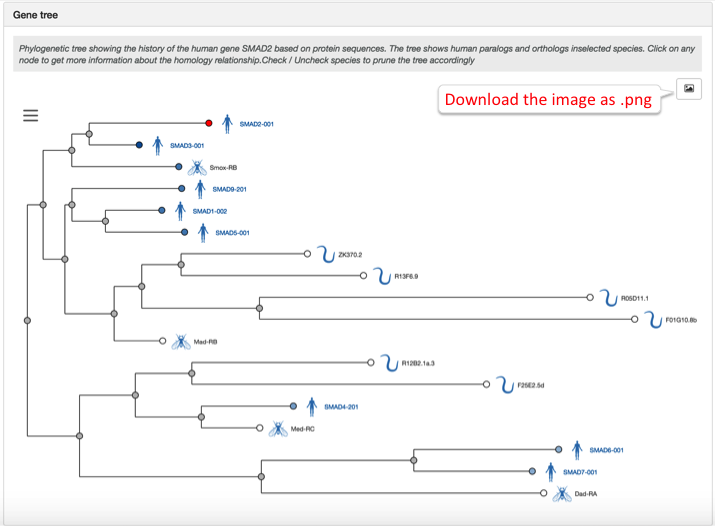
In addition, you can now find targets in human by searching for their orthologues in fly (D. melanogaster) and worm (C. elegans). For example, if you search for the fly dyb gene, you will find its human orthologue, DTNA.
Export Widgets and Share URLs
You can download our widgets as a .png file, with scale factors ranging from 1x (lower resolution) to 3x (higher resolution), and use them in your slides and publications.
Also, the 'Bubbles', 'Table' and 'Tree' views in the Diseases associated with a target pages are now saved in their URLs:
https://www.targetvalidation.org/target/ENSG00000157764/associations?view=t:bubbles
https://www.targetvalidation.org/target/ENSG00000157764/associations?view=t:table
When you share the Table view with your colleagues, they will see the view you have selected. You can also share searches and much more.
Interactive Showcase of Text Mining Results
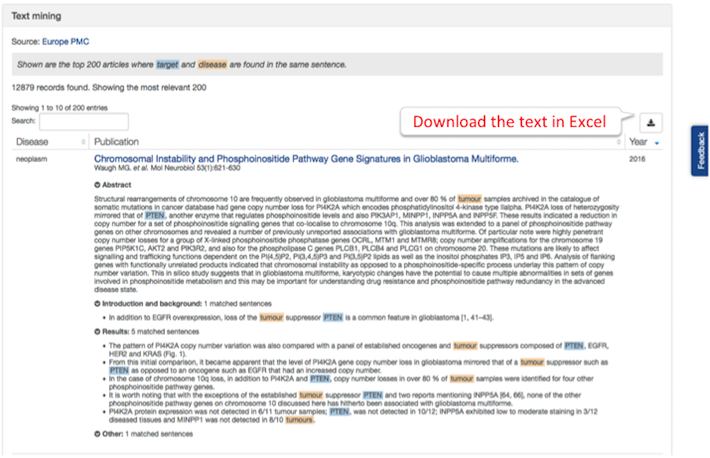
You can see whether the matched entities (i.e. target and disease) are highlighted in the 'Abstract', 'Introduction' or elsewhere in papers resulting from the EuropePMC text mining. The showcase of these sentences is now more interactive and user friendly.
R Client for the Open Targets REST API
If you know R, you can query our API and download all target-disease associations into convenient data frames with the R client package:
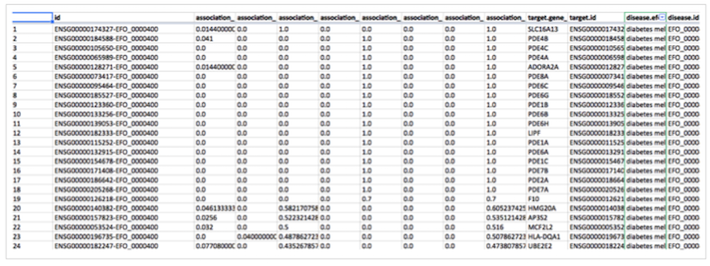
The package and the source code for ropentargets are available on GitHub.
Check our post on How to access Open Targets with R for more details on ropentargets.
Extra Channels for Help and Support
It's now much easier to get help and support with the 'Feedback' and 'Follow us' buttons on the homepage. When exploring our Platform, if something does not look right to you, use the 'Feedback' button on that page and let us know what the problem is.
You can check our updated help pages too, a great place to find out more about the Target Validation Platform:
- Getting started
- Data sources
- Association score
- Frequentely Asked Questions
- Open Targets Platform REST API
Latest Version of Elasticsearch
Our software architecture is more scalable and future-proof with our move to Elasticsearch 2.3, based on Lucene 5.5.0.
Do email us us if you have suggestions or comments on the Target Validation Platform v1.2.