Open Targets Platform 26.03 has been released!
The latest release of the Platform—26.03—is now available at platform.opentargets.org.
Key points
This release includes:
- A new pipeline to enhance the extraction of clinical data—and we have redesigned the widgets displaying this data across our pages.
- Enhancer-to-Gene (E2G) predictive features embedded in our L2G pipeline, improving predictions.
- New data from the GWAS Catalog, and regular updates from other sources including IntAct and String.
- A number of technical updates.
Also, Open Targets Platform data is now available on AWS through the Open Data Program.
For a full list of updates, take a look at the release notes. For a list of key stats and metrics, see the Open Targets Community.
A new clinical mining pipeline
Clinical trials provide key data to the Platform, feeding into our entity profile pages and informing target-disease associations and target prioritisation.
We have implemented a new clinical mining pipeline which expands the data sources for clinical information. Our revamped pipeline fetches, processes, and annotates data from:
- ClinicalTrials.gov via AACT
- ChEMBL curated indications
- ChEMBL drug warnings
- Therapeutic Target Database (TTD)
- European Medical Agency (EMA) Human Drugs
- Japan's Pharmaceuticals and Medical Devices Agency (PMDA) approvals
Some of these aggregate data from other sources.
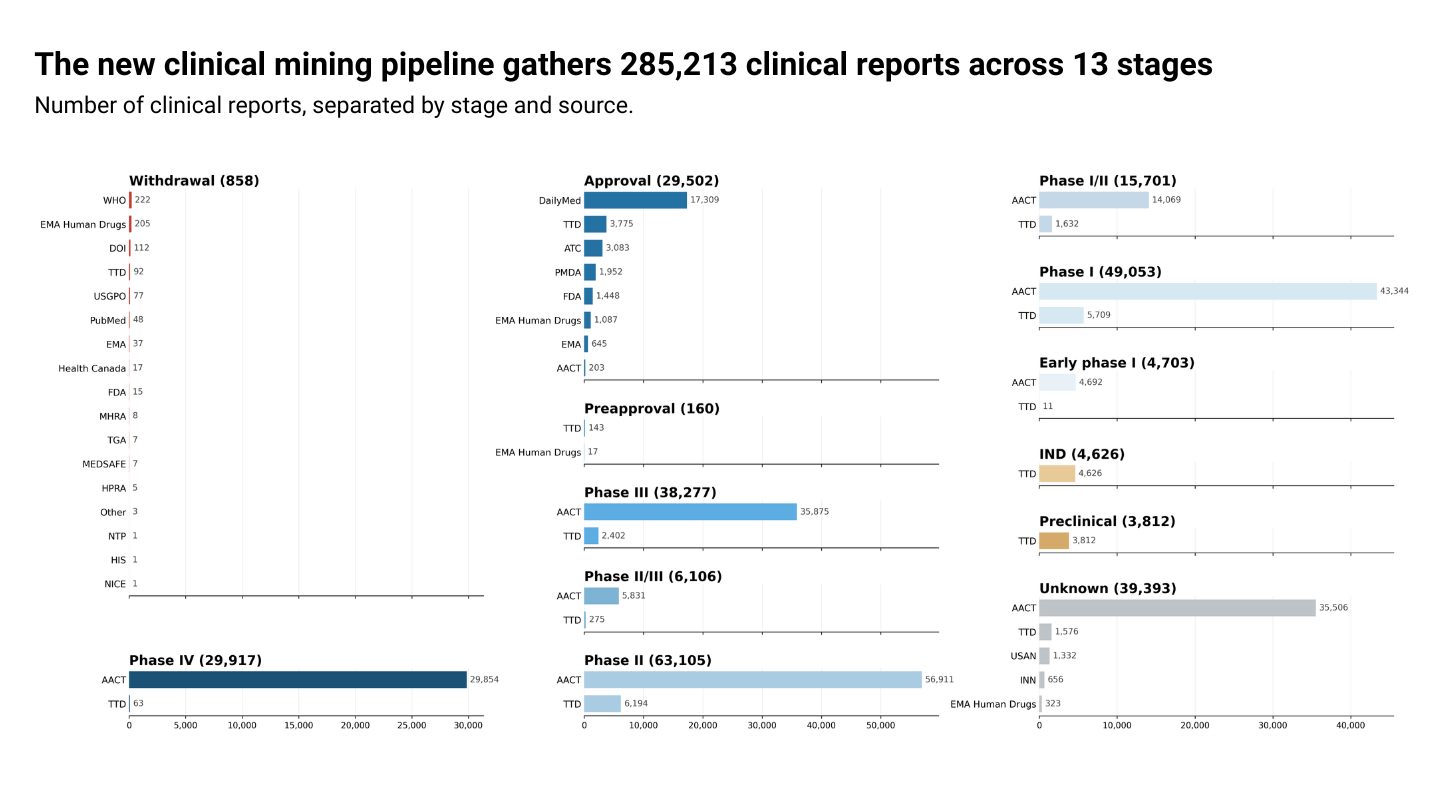
By accessing clinical reports directly, we now have more granular clinical stage information. The pipeline is also more automated and adaptable, which will allow us to iterate faster in the future. This is the first of many changes we have planned to enhance our clinical annotations.
Find out more about the pipeline in the documentation.
Indications (drug pages), Drugs and Clinical Candidates (target and disease pages, target prioritisation)
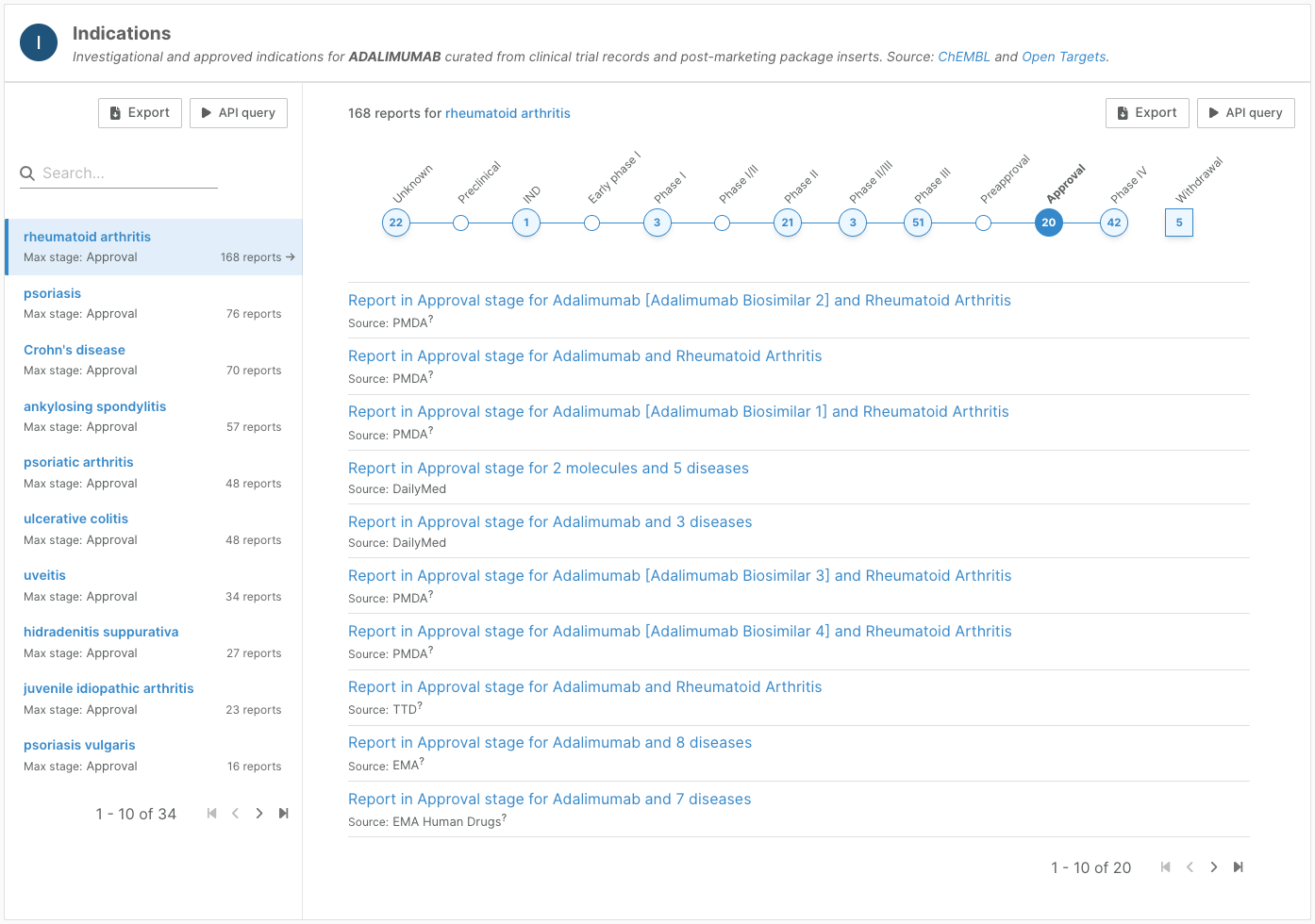
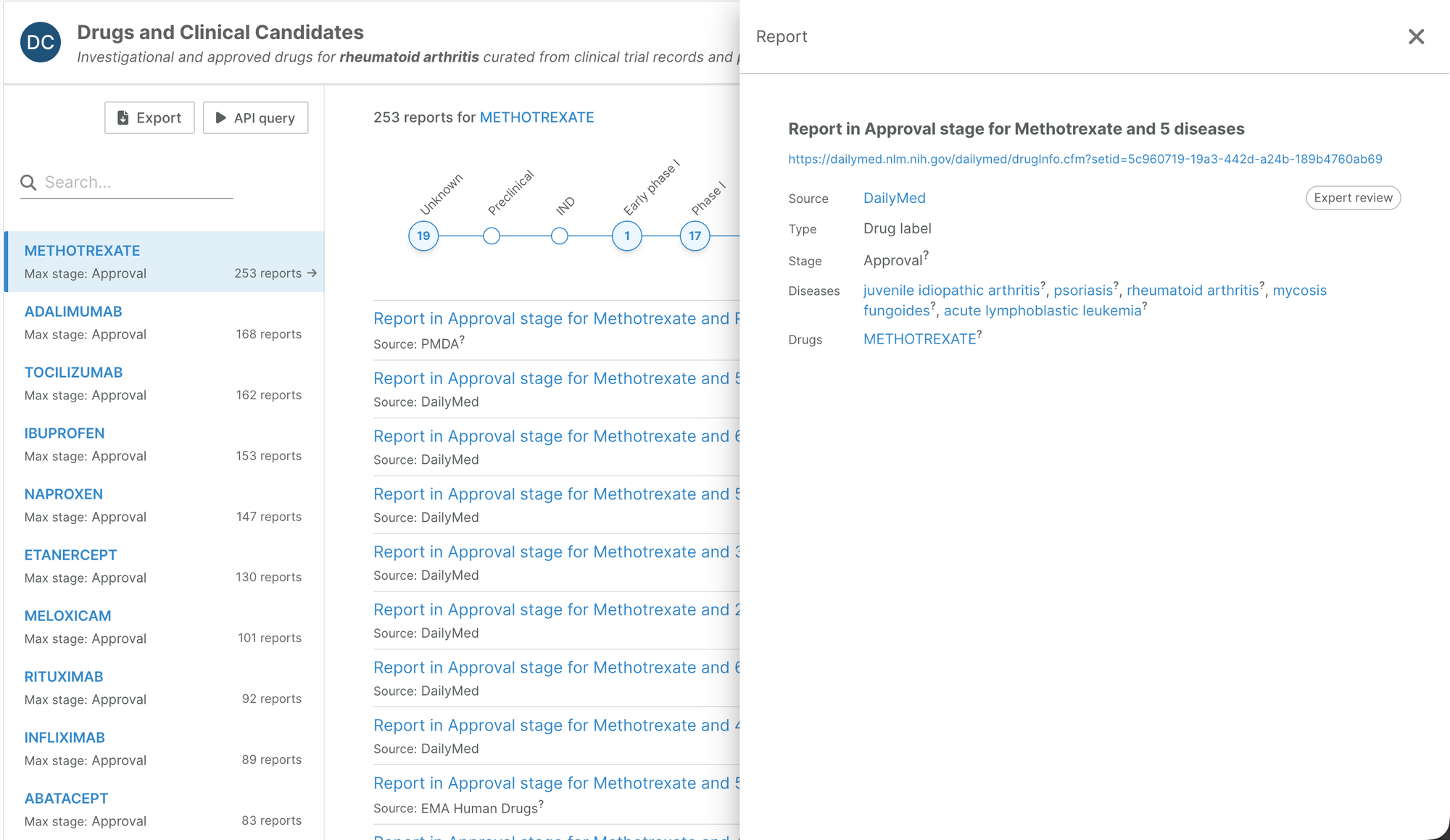
On entity profile pages, we have redesigned the indications widget to aggregate reports by indication (drug pages) or drug (target and disease pages, target prioritisation).
The widget now clearly shows for each indication whether the drug has been approved, and if not, the highest clinical stage reached. We have also been able to add additional nuance to the clinical stages.
Each report can then be explored in more detail.
Clinical Precedence (associations pages)
We have also streamlined the widget providing clinical evidence in our associations pages.
In this release, 102,000 clinical reports contribute to target-disease evidence for 2,592 diseases. This includes 410 more diseases than in the previous release.
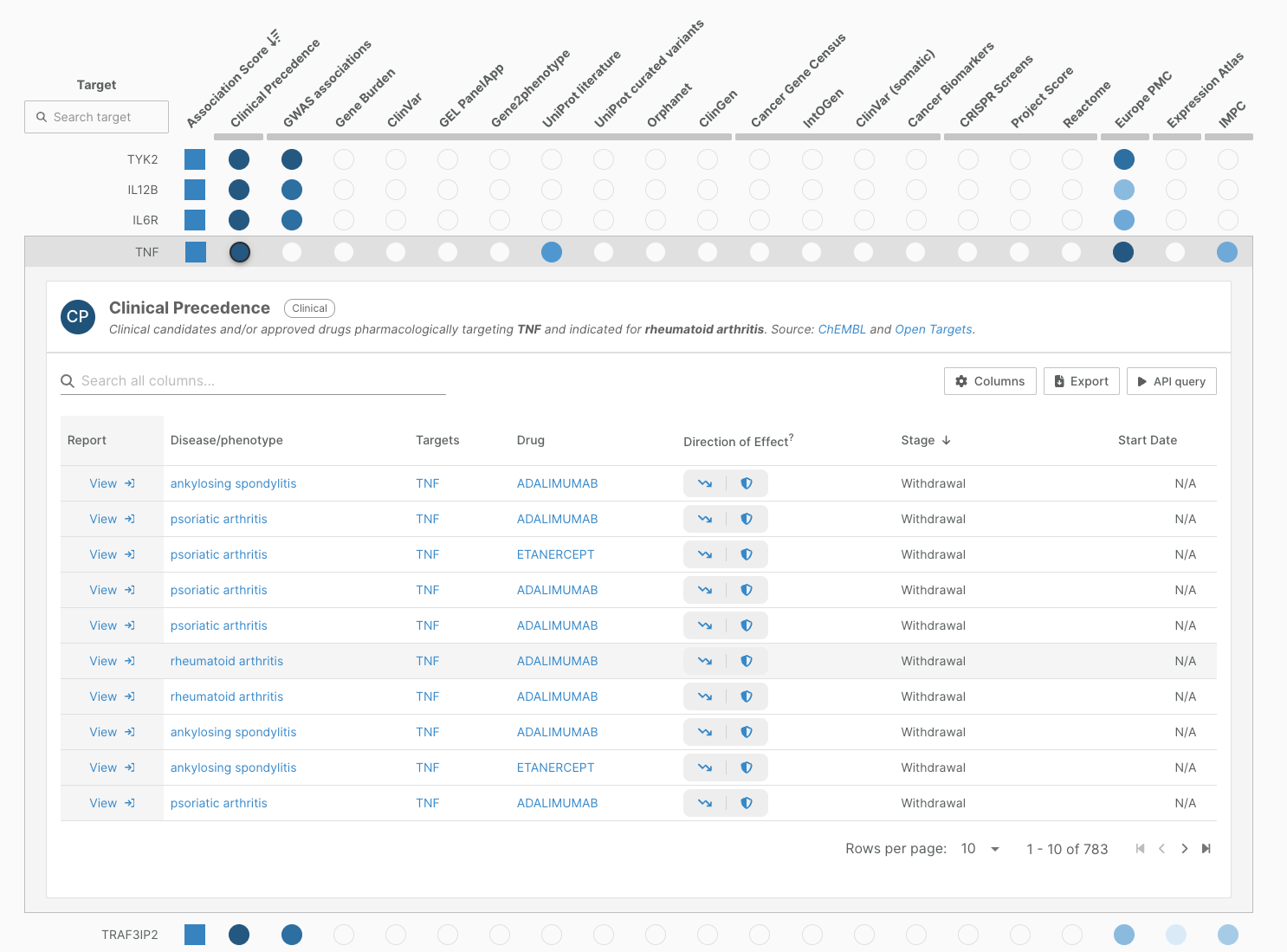
Note that the widget has a new name: Clinical Precedence, better reflecting the fact that the evidence includes reports from therapies that have not yet been approved.
We have also moved this column into the first position of the associations view, as user feedback indicated this was a key starting point in prioritisation workflows for understanding whether a target or an indication is or has ever been investigated clinically.
Enhancer-to-Gene predictive features added to L2G
Last year, we ingested the full dataset of predicted enhancer-gene regulatory interactions in the human genome from the ENCODE-rE2G model.
ENCORE-rE2G integrates molecular features and large-scale perturbations to predict links between variants and transcriptional regulatory elements in a given cell. Each element-gene pair is scored based on the probability of the regulatory effect.
We have used these scores to add two features to our Locus-to-Gene (L2G) machine learning model: e2gMean and e2gNeighbourhoodMean, adding regulatory context to L2G features.
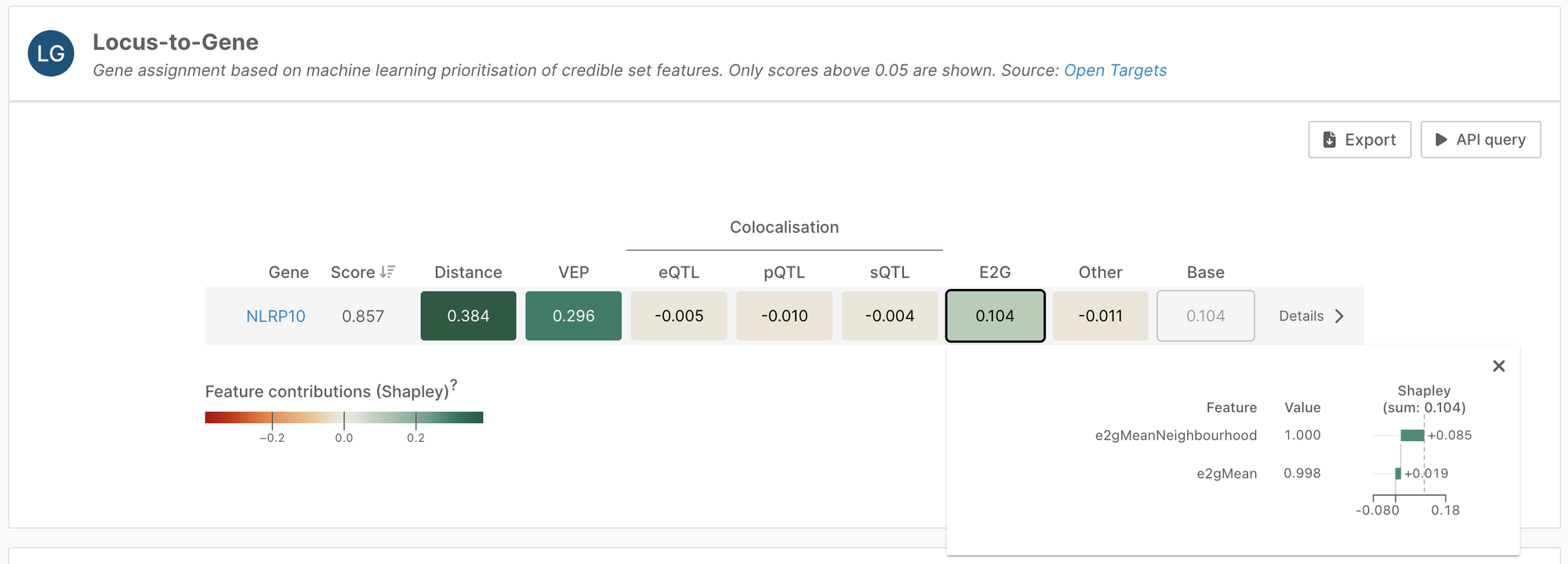
This addition increased the selectivity of the L2G model by 5%; we now have a higher proportion of credible sets with only one assigned gene.
Find out more in the documentation.
Data updates
New GWAS Catalog studies
We have integrated 710 new studies from 97 publications added to the GWAS Catalog since our last release, resulting in over 5,000 new credible sets.
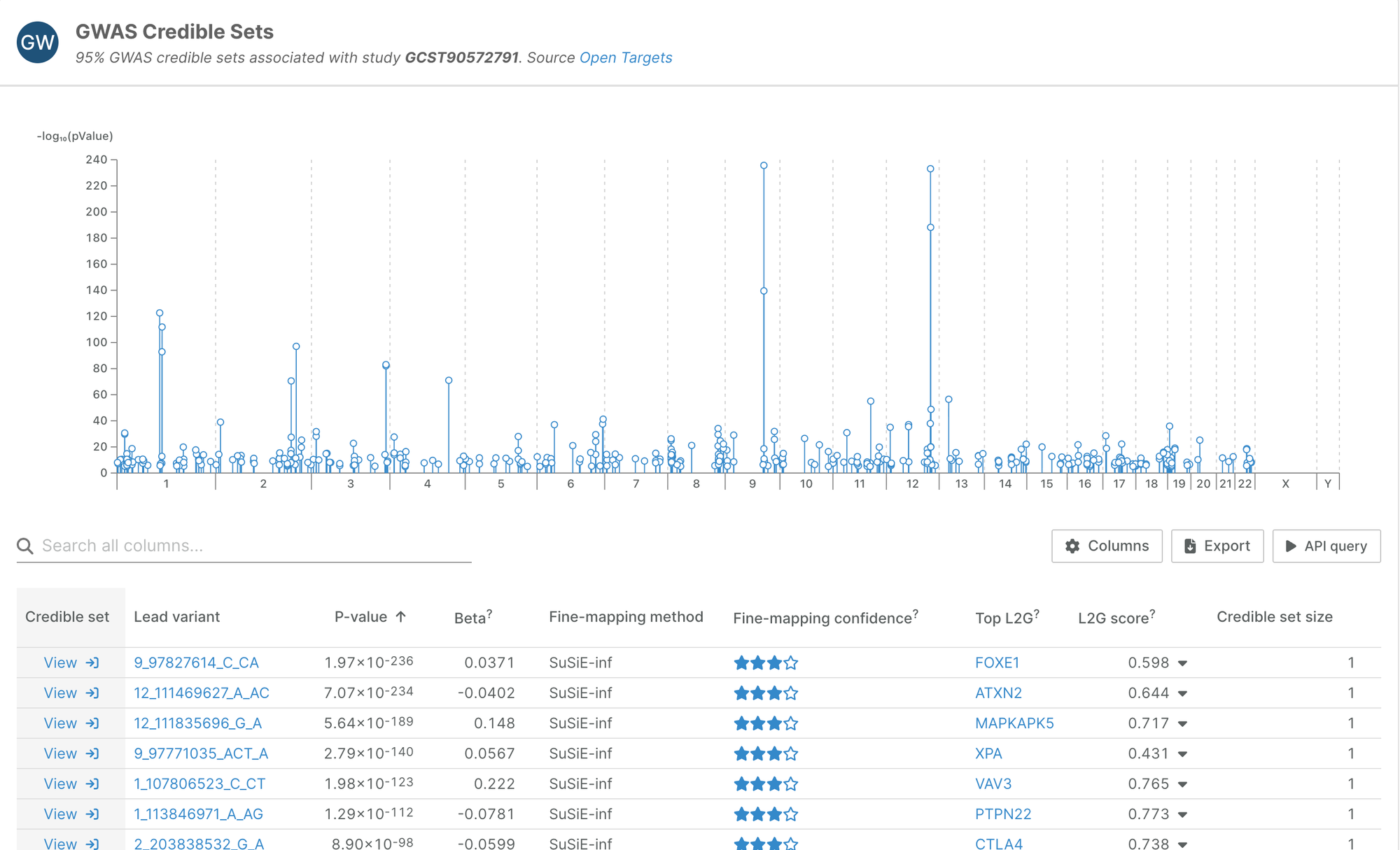
758 of those credible sets come from three studies in a single publication; a genome-wide meta-analysis of hypothyroidism, free thyroxine, and thyroid-stimulating hormone (Rand SA et al. Nat Genet, 2025).
This is the biggest study for hypothyroidism we have ingested, and the first with summary statistics, which means we were able to fine-map them with the more accurate SuSiE method.
Other updates
This release features updated data from String and IntAct, and updates from ClinVar and ClinPGx through the European Variation Archive.
Technical updates
The team has been working to improve the Platform in the background. We have made a number of technical updates and fixes, including:
- Resolving disease/phenotype duplications in the ontology: Users reported on the Open Targets Community that terms could sometimes be represented in the Platform as both a disease and a phenotype. Where this occurs, we have resolved the issue by merging the evidence for the two terms, resulting in the removal of around 300 duplicates.
- Fixed minor scoring inconsistencies between the API and the web interface: a user reported a difference between results given by the API and by the web interface when querying associations. This was due to a discrepancy between the default settings of the API and interface, which we have now resolved.
- Update to the LD annotation: We have fixed the liftover process that gave us incorrect coordinates and impacted the credible sets fine-mapped using GnomAD v2.1 LD hail table (fewer than 1%). Find out more in the issue.
- Clickhouse migration: All the non-search datasets that were still in OpenSearch are now living in ClickHouse. That has more than doubled the speed in most queries, and greatly improved reliability.
- End to end testing: The Front End team implemented End-to-End testing to improve the stability of the Platform. Test suites can now be run on configured test pages of interest. Find out more in the project readme.
Data access: AWS and MCP
Open Targets Platform data is now available on Amazon Web Services (AWS) through the Open Data Program.
We have also released an update to the official Open Targets MCP. Notably, the MCP tries to first reduce the context to the domain of interest, improving performance on Q&A and significantly reducing token usage.
Find out more about the MCP, and all our data access options, in the documentation.
As usual, please share any comments, questions, or suggestions on the Open Targets Community.