Open Targets Platform 23.12 has been released!
The latest release of the Platform — 23.12 — is now available at platform.opentargets.org.
Key points
In addition to our regular data updates:
- This release introduces a completely new view: Target Prioritisation. It provides an assessment of the target features considered when prioritising (or deprioritising) targets for drug discovery
- The Platform now pulls in pharmacogenetics data from PharmGKB through the European Variation Archive
- We have added new features to our Associations on the Fly view, such as allowing users to download customised entity lists and to pin targets of interest
Key stats
| Metric | Count |
|---|---|
| Targets | 62,733 |
| Diseases | 25,246 |
| Drugs | 17,095 |
| Evidence | 16,710,896 |
| Associations | 7,994,180 |
Additional metrics are available on the Open Targets Community.
Target prioritisation
A major new feature of this release is the Target Prioritisation view, which provides an assessment of target-specific properties of interest when evaluating the suitability of a target for a drug discovery programme.
The new page can be accessed from the target-disease associations page when searching for targets associated with a disease, and clicking on the target prioritisation factors tab. With a similar structure to our associations-on-the-fly view, targets are sorted by their overall association score for the disease of interest, and users can browse a series of target annotations directly in the view.
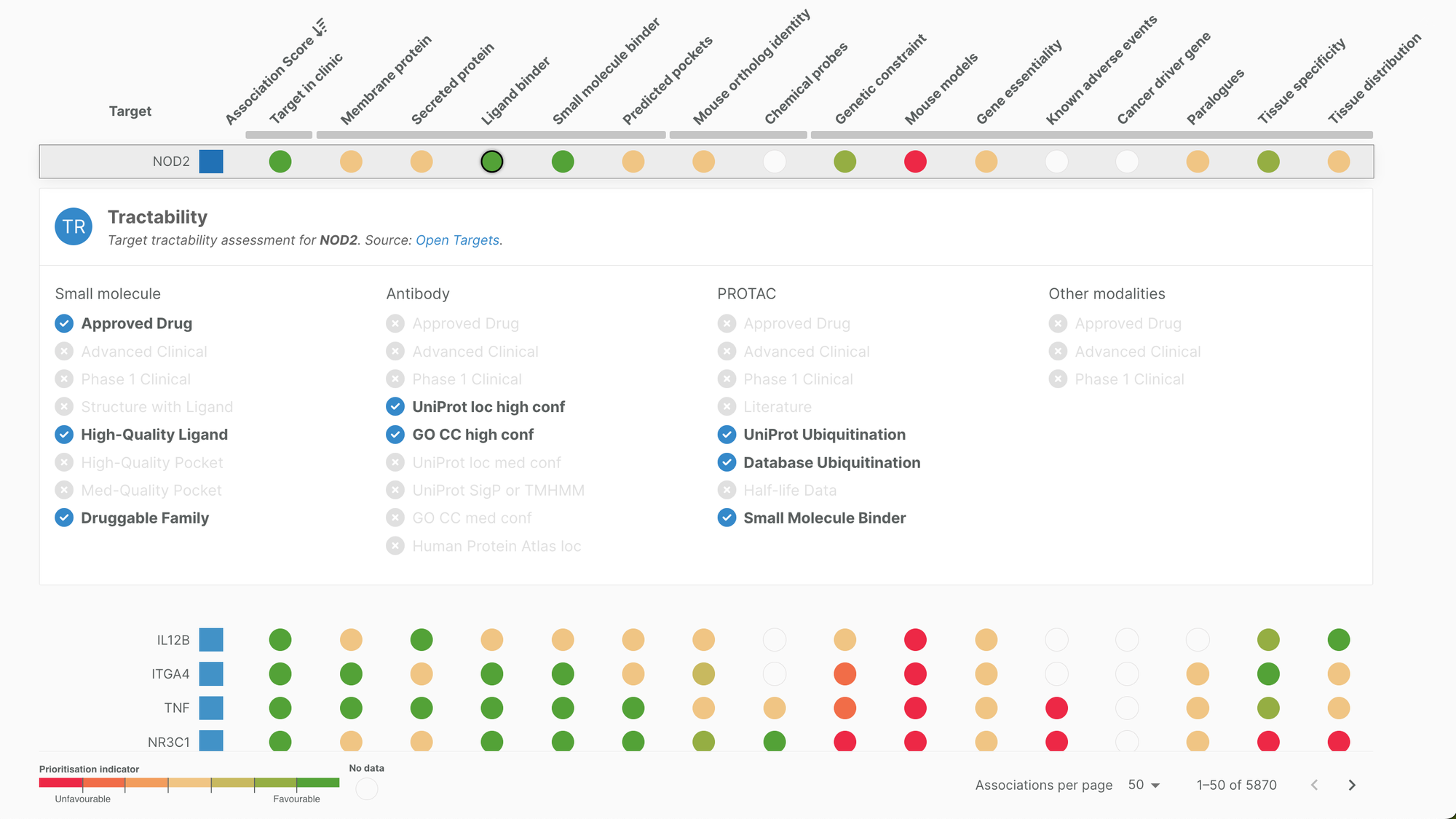
The data showcased in this view is also available in our target profile pages. It is presented here with an additional layer of analysis with a drug discovery focus. We have designed a traffic light scoring system to provide users with an at-a-glance overview of target attributes. On this colour scale, green indicates potentially positive attributes while red indicates potentially negative attributes that may contribute to a target’s prioritisation or deprioritisation, respectively.
Target properties have been aggregated into four categories — Precedence, Tractability, Doability, Safety — and individually scored as part of an Open Targets project. Find out more about the scoring in the Platform documentation.
These assessments will evolve as part of the project progression, so any feedback regarding this view is very welcome.
Introducing pharmacogenetics data to the Platform
This release introduces a new datasource to the Platform: the pharmacogenetics widget will incorporate data about the impact of human genetic variation on drug responses. You can find this data on the target and drug profile pages.
The new widget showcases relationships between drugs, diseases/phenotypes, and genes, including details on genetic variants that influence drug metabolism, efficacy, and adverse effects.
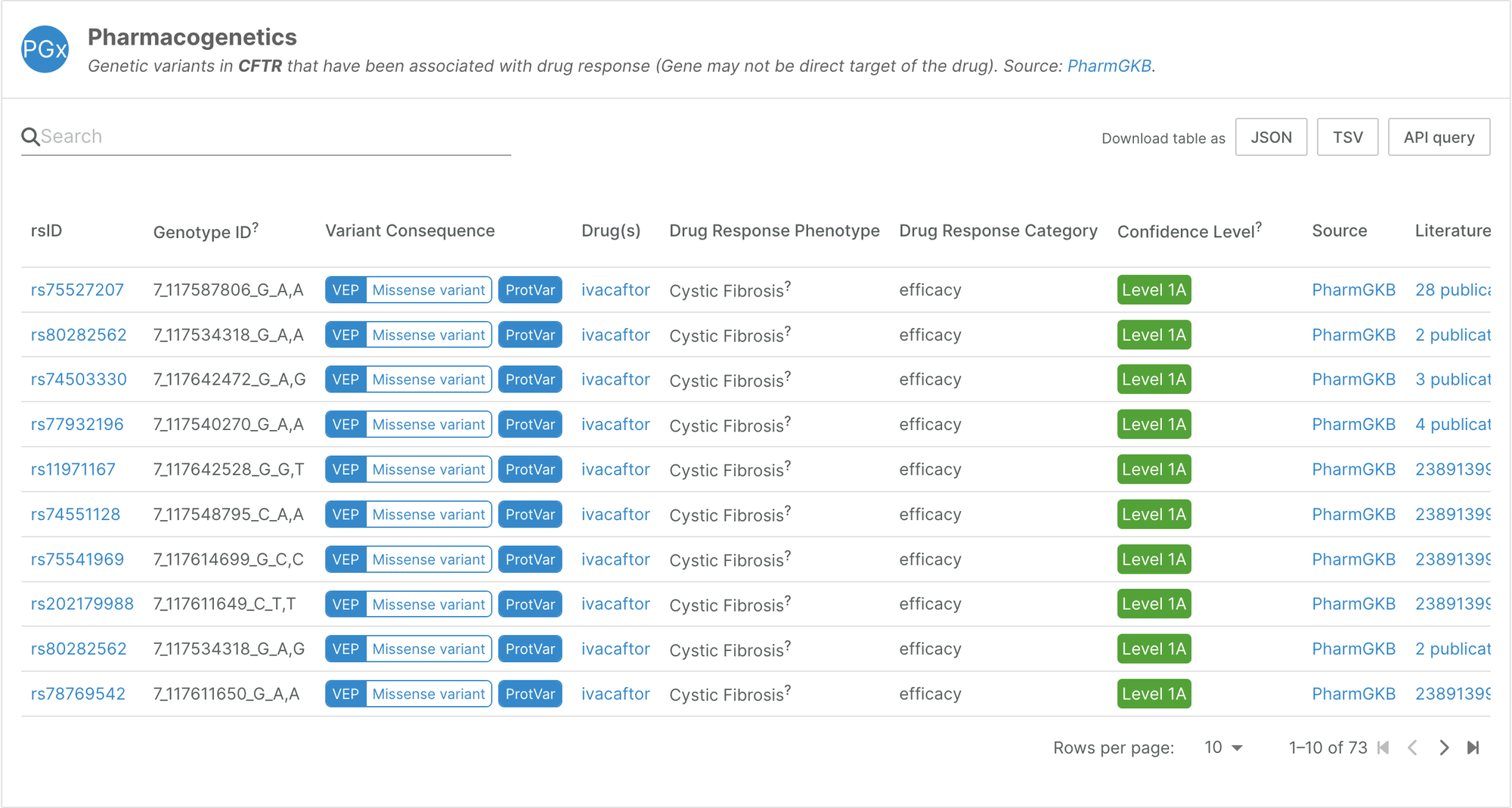
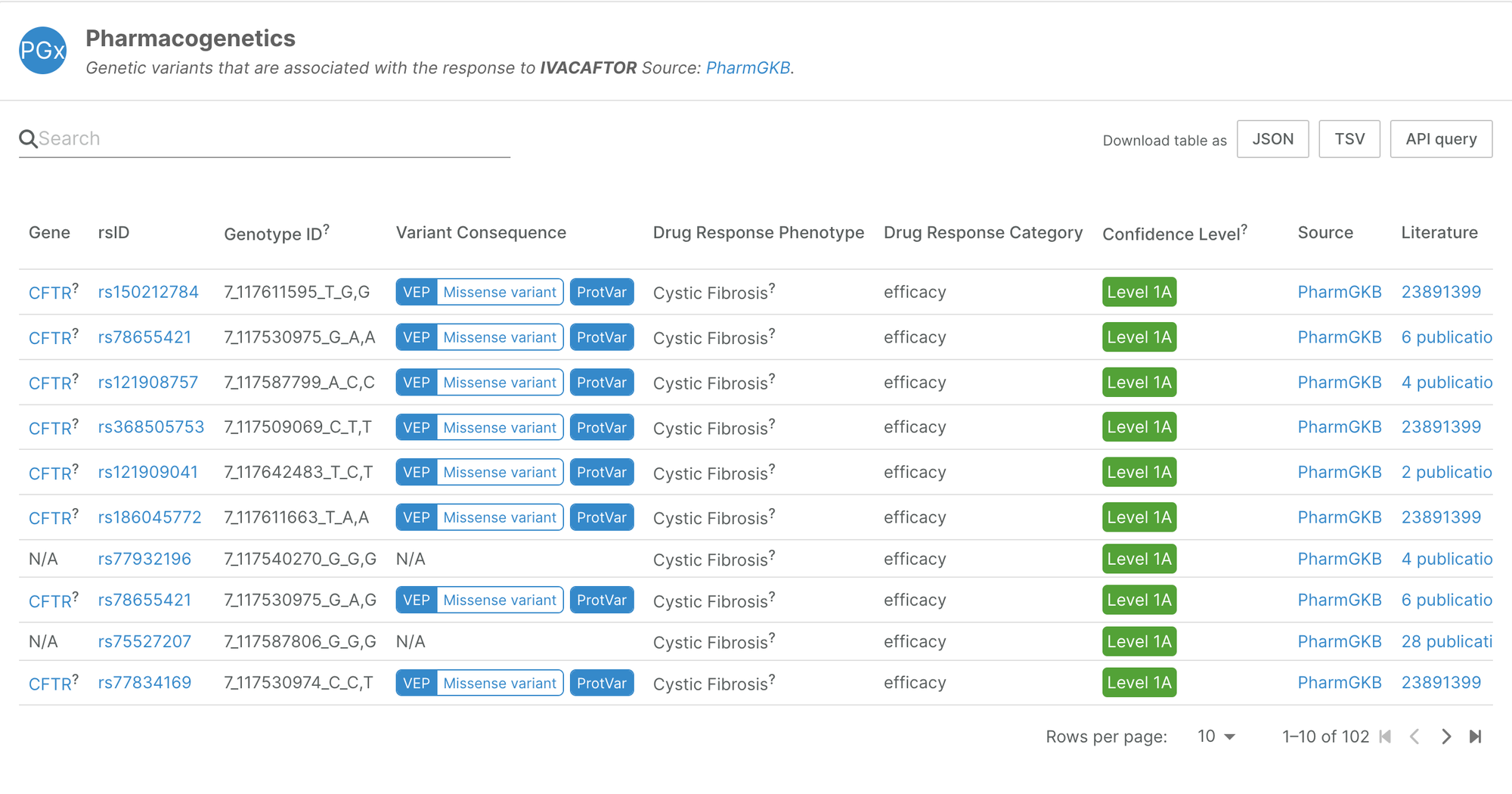
The first release of this datasource integrates data from PharmGKB, a pharmacogenomics knowledge resource. Amongst other activities, the PharmGKB team annotates genetic variants and gene-drug-disease relationships, and summarises important pharmacogenomic genes, associations between genetic variants and drugs, and drug pathways.
This provides information on genetic variants that affect a patient’s response to a drug, allowing additional granularity to therapeutic hypothesis formulation, taking into account how specific genetic backgrounds can influence treatment outcomes.
The Platform only showcases records of high confidence (1A, 1B, 2A and 2B in PharmGKB’s categorisation system). In future releases, we will focus on streamlining the integration of variants reported as star alleles, ensuring our pharmacogenetics data aligns with the common notation used by clinicians and researchers in the field.
Improvements to Associations-on-the-fly
Thanks to feedback from our community, we have been working on improvements to the associations view we introduced in the previous release (23.09).
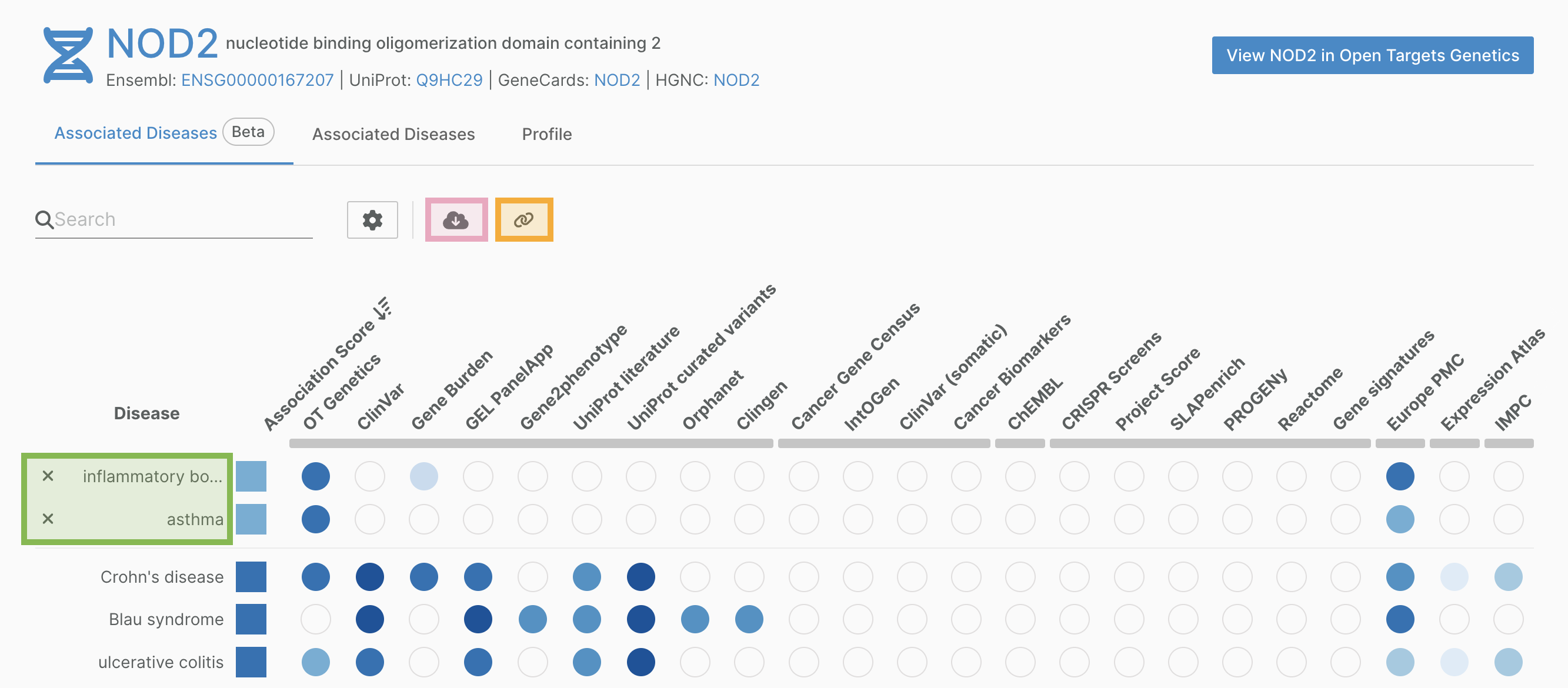
Associations-on-the-fly now allows users to export the associations table and the target prioritisation table in json or tsv format. The export parameters can be customised to the user’s preferences.
We have also added the ability to pin rows of interest. Pinning rows creates a customised, shareable url for easy reference.
This was the last release of the year — but we have more in store for you next year! As usual, please share any comments, questions, or suggestions on the Open Targets Community.




