Open Targets Platform 20.06 has been released!
We are super thrilled to announce that we have a new release of the Open Targets Platform.
Release 20.06 is full of exciting new evidence and updates on data sources as well as tweaks to the target tractability pipeline and to data from Open Targets Genetics.
And on top of all of this we have been working on a brand new design of our user interface, with an alpha version now available for you to have a peak at.
First things first, let’s check our favourite stats for this new release:
| Targets | Diseases | Evidence | Associations |
| 27,596 | 13,914 | 10,154,924 | 7,282,832 |
Looking for a breakdown of these numbers per data type and data source? Head to release notes or use the REST API stats endpoint.
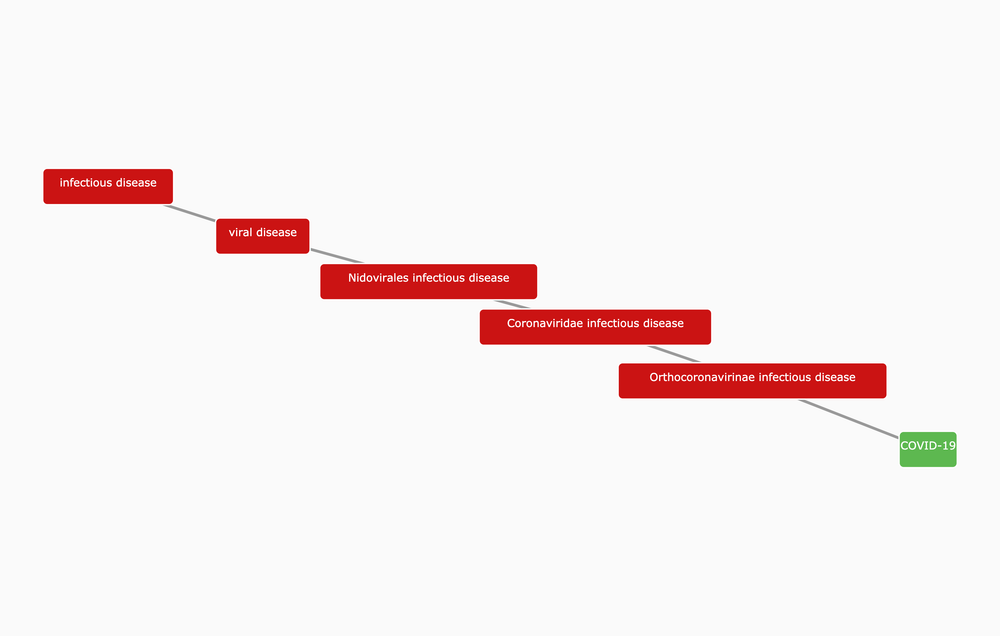
COVID-19
We are excited to announce that COVID-19 is now available in the Open Targets Platform.
In this release we integrate drug information from 758 clinical trials and 1,228 evidence strings from the literature to identify 261 targets associated with COVID-19.
Besides COVID-19, other 95 diseases and phenotypes are also included for the first time in the Open Targets Platform. Huge thanks to the ongoing efforts of our dedicated collaborators for curating these new disease terms!
Check Open Targets and COVID-19 to recap on what else you can do with our data to identify and prioritise drug targets for the treatment of this new and devastating disease.
Open Targets Genetics evidence
Since the integration of Open Targets Genetics evidence as a new data source for genetic associations in the Open Targets Platform, we have noticed that a huge number of common genetic variants had a very low score: a whopping 1.65 million evidence strings had a score between 0.00 and 0.10!
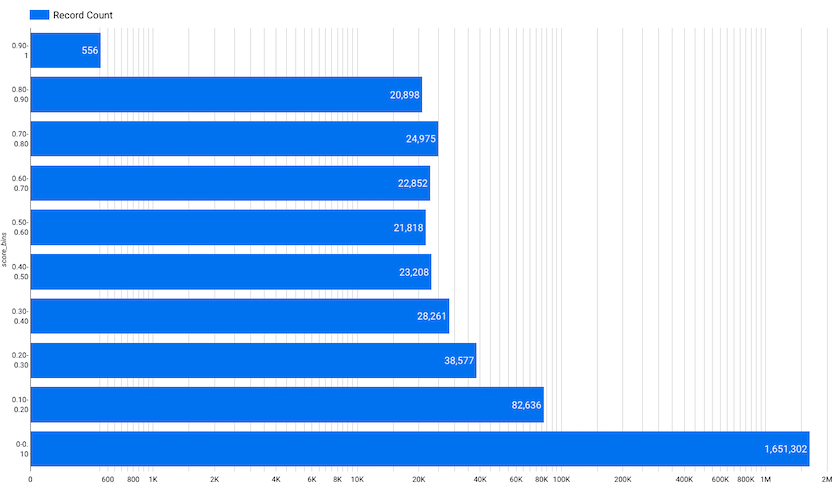
Record counts per score bins for Open Targets Genetics evidence available in release 20.04 of the Open Targets Platform. Image credit: Asier Gonzalez
For release 20.06, we have applied a threshold to this evidence and now we only use common variants with locus-to-gene score equal to or greater than 0.05.
By removing low-scoring evidence, we ensure the remaining data is more relevant and accurate, so that you can be more confident in the target-disease associations supported by common variants in the Open Targets Platform.
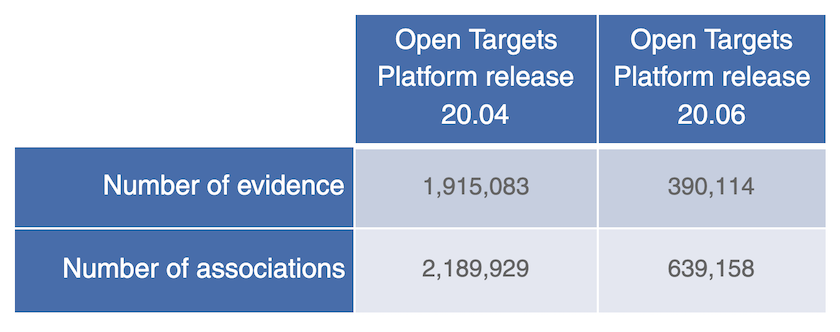
Differences in the number of evidence strings from Open Targets Genetics and the associations supported by this data in previous and current release of the Open Targets Platform.
We will continue to monitor this in light of our commitment to integrating robust human genetics and genomics data for systematic drug target identification and prioritisation.
Target tractability pipeline
We have implemented minor changes and fixes to target tractability (also known as druggability) for two of its drug modalities, small molecule and antibody.
Small molecule
Up to release 20.04, the max_phase for the small molecule tractability assessment was the same as the max_phase reported for its target. We have changed this to take into account the max_phase for the drug compound itself.
Confused? Let's look at an example to illustrate this.
ITGA4 has one small molecule in phase 3 (firategrast) and two antibodies in phase 4 (natalizumab and vedolizumab). Therefore the max-phase for ITGA4 was set to 4 and the tractability for Small molecule for ITGA4 was assigned to "Clinical precedence | Phase 4". However, there is no small molecule in phase 4 but rather in phase 3 targeting ITGA4. So the max_phase for firategrast is now used and the assignment amended to "Clinical precedence | Phase 2 or 3".
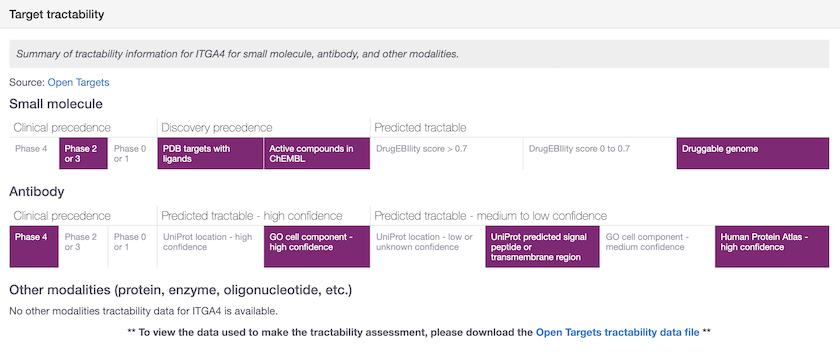
New bucket assignment for Small molecule following changes to which maxphase is used in the tractability pipeline.
Antibody
We now include new terms for UniProt subcellular locations that were missing in previous releases e.g. "Peripheral membrane protein" in ACY3, and also include the latest changes to the "Human Protein Atlas - high confidence" bucket following modifications to their reliability score.
We will continue to work with our ChEMBL colleagues to fine-tune the target tractability assessment with more new features coming up in a few months' time.
Other news
Additional exciting news for release 20.06 is listed below:
-
Almost 20,000 extra PheWAS evidence strings revealing novel associations e.g. WNT3 in muscle strain
-
Brand new chemical probes from SGC and Chemical Probes e.g. glyburide to target ABCC8
-
Most up-to-date differential and baseline expression data from Expression Atlas release 35
New user interface
Did you know that we have been working around the clock on a new, much sleeker and easier-to-use graphical user interface?
We are extremely happy to announce the alpha release of the Open Targets Platform is now live!
Why not start delving into this new interface straight away?
You can start your journey by searching for a:
-
disease or phenotype e.g. multiple sclerosis and paralysis
-
drug e.g. hydroxychloroquine and adalimumab
-
target e.g. EGFR
We hope you enjoy exploring release 20.06 of the Open Targets Platform (and its alpha version) as much as we have enjoyed working on this data in the last couple of months.
And as usual, get in touch if you want to discuss any of these new features or if you have suggestions for the team.