Next steps in our web applications' architecture
My name is Carlos Cruz. I'm a software engineer focused on front end. As part of the Open Targets team, I work on maintaining and developing the Open Targets web applications.
At Open Targets, we maintain two UI applications: the Open Targets Platform and Open Targets Genetics. Both are JavaScript applications that use React, GraphQL, and MUI, among other libraries. They have given us good results in terms of performance and development. However, we are bumping against the limitations of sharing components between the two applications and maintaining a foundation of good development practices (documentation, code consistency, testing).
Our next Open Targets Platform release will introduce a monorepo for our front end. Here, we explore why we made this change, and our next steps.
A brief history of our Front-end ecosystem
We launched the Open Targets Platform at the end of 2015. Initially, the front-end consisted of an Angular v1 application. Three years later, we launched Open Targets Genetics. From a software perspective, Genetics was a proof of concept, refactoring the Platform code using React. Given its success and versatility, we incorporated React into our Front-end ecosystem.
We rewrote the Platform’s UI basing ourselves on the Genetics code. We, therefore, developed OT-UI, an intermediary library to maintain a source of truth between the components of both user interfaces. In my opinion, it was the best way to solve the problem of sharing features. And it worked. With the help of OT-UI and based on Genetics, we successfully created the Platform, and both applications shared similar architecture and components.
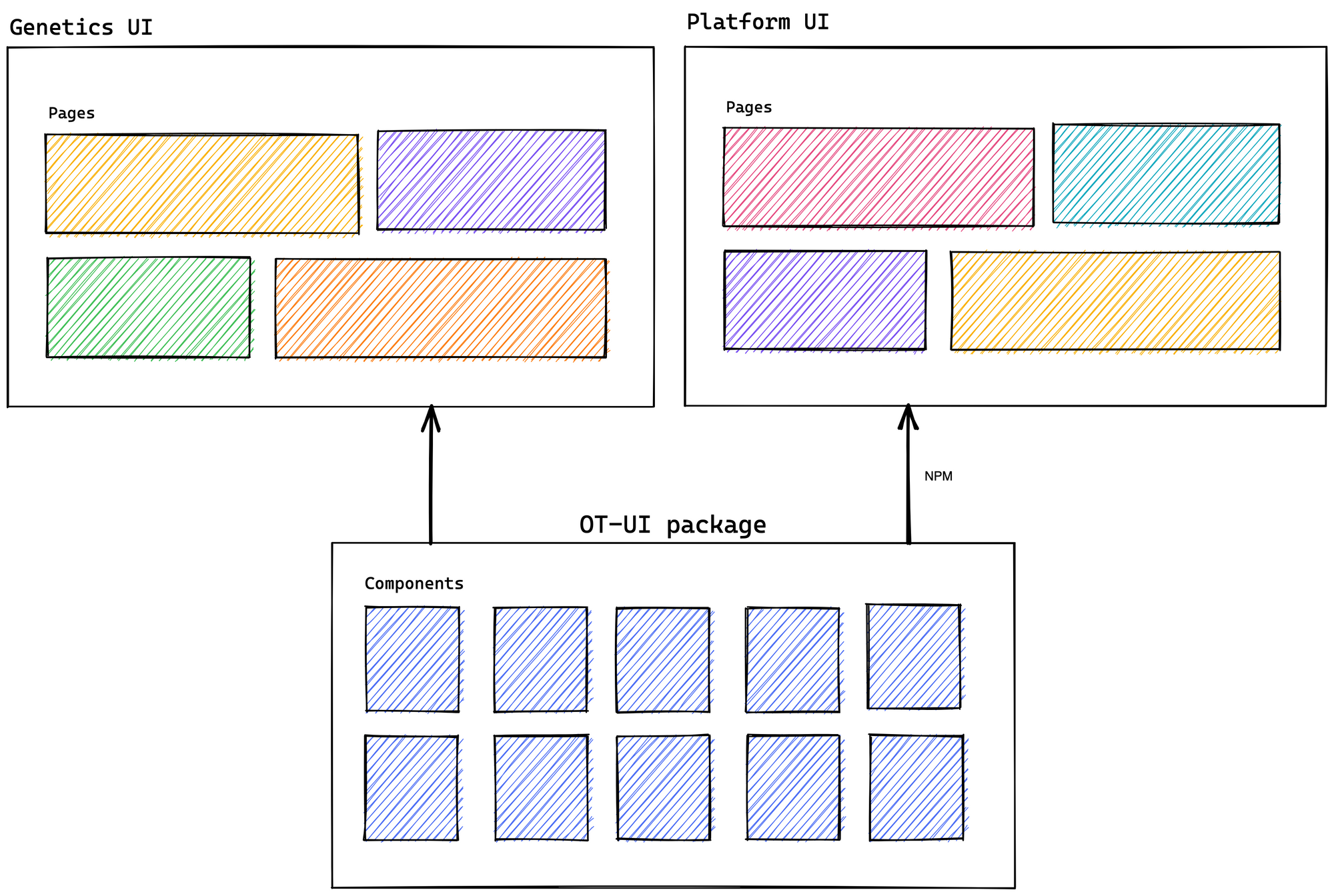
From the theory to the practice
After this first implementation, we started running into difficulties working in an agile way with the component library. The use of OT-UI was complicated by issues such as handling third-party library versions, developing in a local environment using NPM (node package manager), and integrating custom components. Because of this, we concentrated our development efforts on the Platform UI. While this allowed us to maintain consistency within the Platform, it altered the Platform’s architecture without taking into consideration Genetics or the OT-UI library.
Over time, the gap between the projects widened, and eventually, the Platform stopped using OT-UI entirely, only updating relevant libraries such as Material-UI and GraphQL. The project also implemented a new architecture more in line with the design and user experience needs. On the other hand, Genetics continued to integrate the required components from OT-UI, and this ended our attempt to maintain a library to share parts of the codebases.
OT-UI was deprecated.
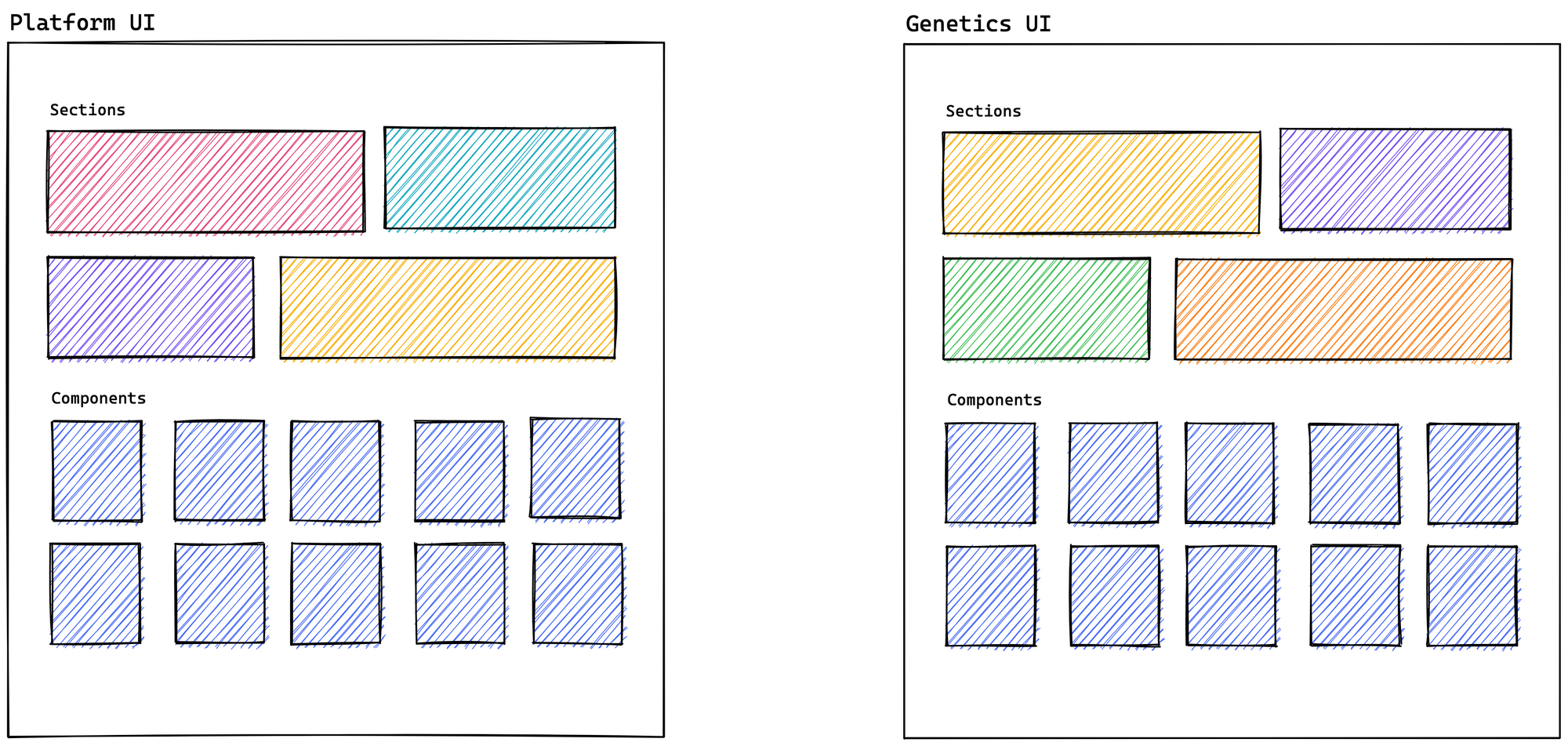
Recently, we have made an effort to update Genetics, and it has given us good results. We are already matching versions of MUI and GraphQL with the Platform, in addition to sharing the same release process. We are even trying to use the layout as it currently exists in the Platform. But this brings us right back to a previous issue: when we need to reuse suitable primitive components from the Platform, technically, we don't have a solution that gives us a desirable development experience.
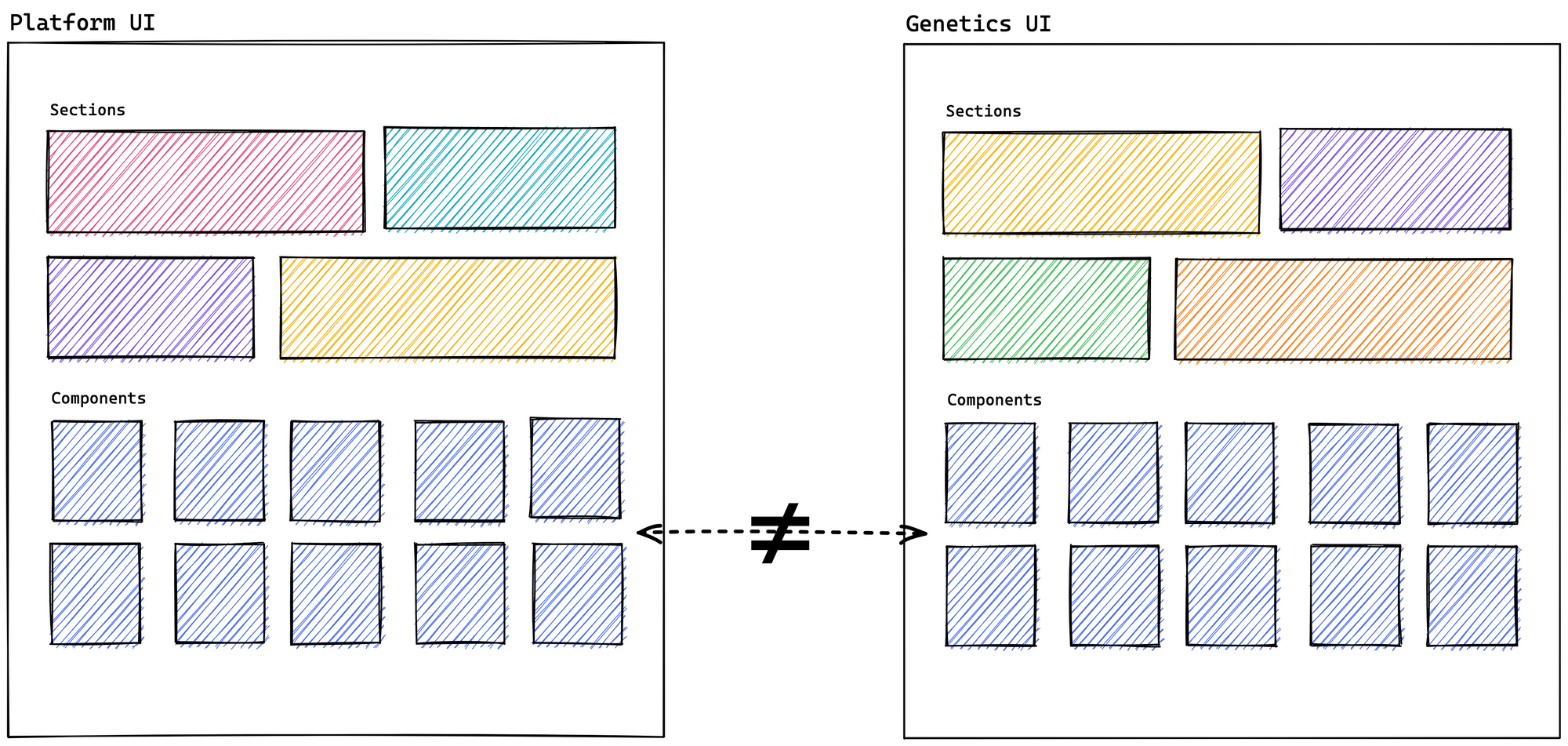
A new solution with Turborepo
After experimenting with different possible solutions and seeking to align our plans for the future concerning the user experience in the user interface, we came up with the idea of creating a monorepo. Both applications will coexist within a single repository and share primitive UI components (BasePage, Header, Chips, Table, among many others). It will even eventually allow us to have the flexibility to easily share entire sections of one application in another with their respective components, query, and test (Literature, Summary, Ontology, and PheWAS, to mention a few).
Among the different options, we explored Lerna, Yarn Workspaces, Git Submodules, Nx, and Turborepo. We found that the closest to our needs that provides greater flexibility and facilitates the development process is Turborepo.
Turborepo will allow us to "move" our applications with the least development effort and have the freedom to update different parts of our architecture gradually. Initially, we will adjust our release system without altering the code of the applications. We will then begin a gradual update of Genetics, removing the remaining OT-UI components and replacing them with shared Platform components. At this point, we are free to explore many other ideas, for example:
- Investigate the use of NextJS in our UIs.
- Create shared visualizations with Svelte or another JavaScript framework
- Improve our implementation of integration tests.
- Integrate complex task pipelines in our codebase.
We will even be able to generate new UIs with our package scheme quickly.
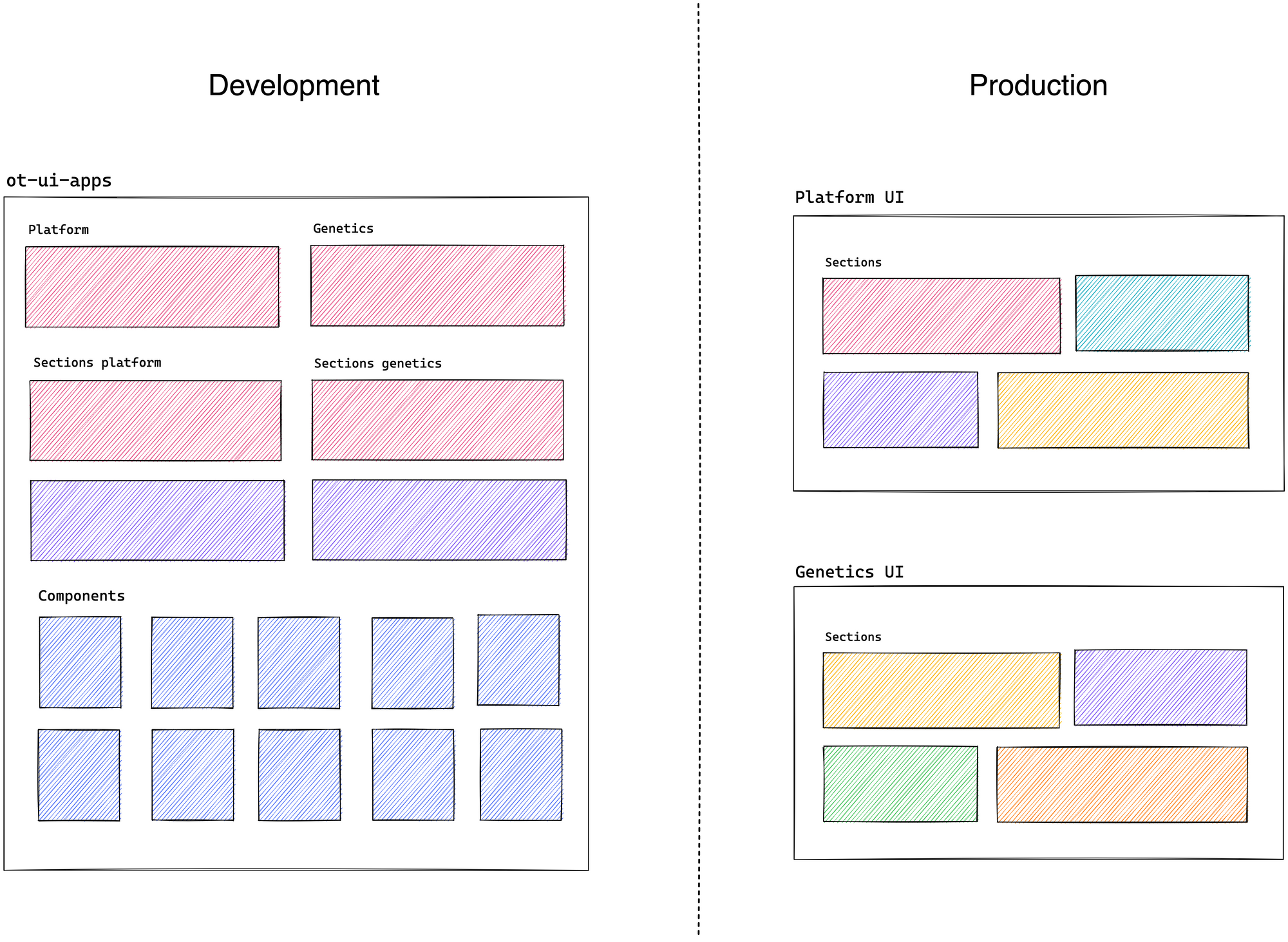
We have started the development and implementation in the OT-UI-APPS repository, and we hope to have advanced for the subsequent releases of 2022.
We would like to hear your opinions and share more details of our experience based on the evolution of our Front-end architecture. I leave attached the GitHub issue, as well as the community post.
Thank you for reading!