Integrating Stem Cell Village Models with the Open Targets Platform for Drug Target Discovery
I have spent the past three months interning in the Open Targets core team. I am currently a PhD candidate in Joseph Powell's lab at the Garvan Institute of Medical Research in Sydney, Australia, where my work focuses on how genetics drives cellular phenotypes in complex diseases. Being a member of the Open Targets team has been immensely fulfilling, and the process of learning and collaboratively applying Open Targets tools to data I collected in the lab has taught me a great deal about translational research.
We each have millions of single-nucleotide polymorphisms, or SNPs, across our genome. Some of these SNPs function as expression quantitative trait loci (eQTLs), which regulate gene expression depending on the combination of alleles present. eQTLs may act in specific cell types, at specific developmental stages, or as a disease progresses. In my research, I aim to uncover the genetic regulatory mechanisms underlying development and variation in drug response across individuals. This will enable the identification of novel genetic targets for drug development and support treatment strategies, ultimately improving patient outcomes.
Building a large-scale differentiation dataset: the Stem Cell Village model
Human induced pluripotent stem cells, which may be reprogrammed from a donor’s adult cells and differentiated to any cell type in the body, enable patient-specific disease modeling in vitro. Because of this capacity, large-scale studies employing stem cells are emerging, providing power for population genetics research and clinical trials in a dish.
To study genetic regulation in vitro, the Powell lab employs “stem cell village models”, in which hundreds of stem cell lines collected from different donors are pooled and cultured together. The village model enables us to run hundreds of experiments simultaneously, empowering us to explore how genetic differences may drive patient-specific differences in development or in responses to medicines.
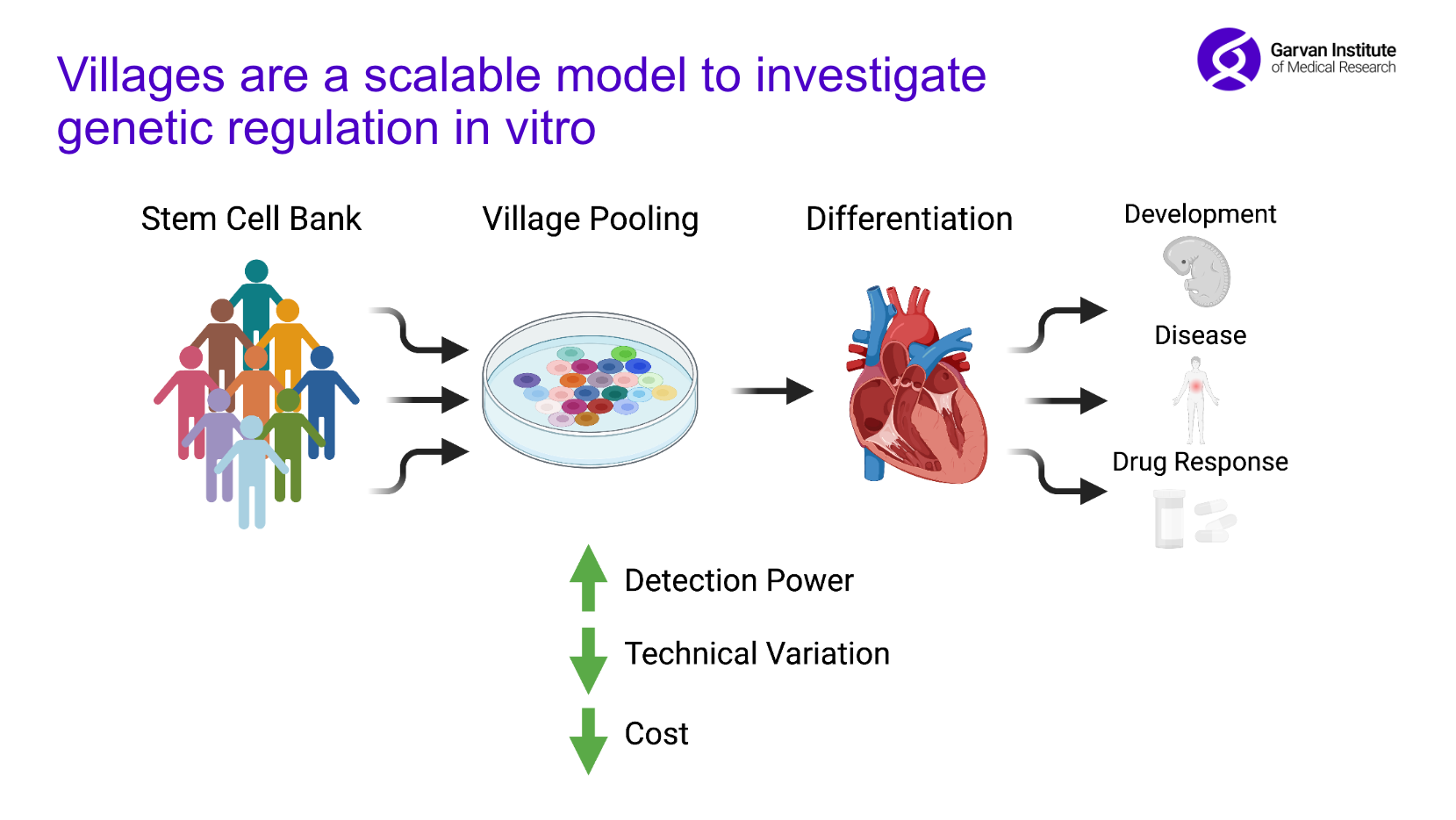
Over my PhD, I generated a large-scale single-cell RNA-sequencing dataset of 2 million cells, in which I pooled stem cells from 150+ donors and differentiated them to cardiomyocytes, a type of heart cell, over 24 days. I collected cells from several time points across this differentiation trajectory, including three critical time windows during which I sampled cells every few hours. This dataset allows us to investigate the fine-grained genetic regulatory mechanisms that act across development and how these mechanisms may differ across individuals.
Connecting gene expression to GWAS signals
From January through March of 2026, I joined the Open Targets team to collaboratively develop and apply translational pipelines to this dataset. Working with Tobi Alegbe (Single Cell Data Scientist) and the broader Open Targets team, we applied colocalisation analysis to determine whether the genetic variants acting as eQTLs in our dataset are the same as those identified by genome wide association studies (GWAS) as being associated with disease. This analysis enables us to identify eQTLs as the missing genetic link between gene expression and disease-associated loci. After running colocalisation, I then assessed whether our colocalisations were enriched for cardiac-related GWAS and, more broadly, how informative my dataset is for studying cardiac disease. Next, I investigated the novelty of my colocalisations and identified several existing cardiac–relevant gene–disease associations for which no genetic data previously existed.
Locus-to-Gene
After exploring relationships between gene expression and disease using colocalisation, I was interested in how adding my data would affect the prediction of causative genes in Open Targets’ Locus-to-Gene algorithm. Locus-to-Gene, or L2G, uses genetic and functional genomics traits to rank the most likely causative genes at each GWAS locus. I found that adding my functional genomics data led to more accurate predictions of causal genes at cardiac-relevant loci.
Drug repurposing
Lastly, the Open Targets Platform contains rich information on genes and the drugs that target them. I used the Platform’s data to nominate candidate drugs for repurposing, spanning both drugs currently in clinical trials and approved medicines.
Now, I aim to validate the findings from my time at Open Targets in follow-up screens, and plan to eventually incorporate my data into the Platform after publication.
The Open Targets Platform presents an unparalleled collection of genomic and therapeutic data, integrating evidence from GWAS, functional genomics, and drug development pipelines to systematically link genes to diseases and potential interventions. Learning about the Platform and its diverse applications directly from the team that built it has been an eye-opening experience. I’m grateful to all the members of the Open Targets team for creating such a rich resource and for hosting me in Cambridge to learn and apply these insights to my data. This experience has shown me how eQTLs can uncover the missing links between gene expression and disease, and has provided me with a framework for evaluating genes in the context of drug development.