Breaking news: extra data in the latest version — 17.04 — of the Open Targets Platform!
We have released our latest version of the Open Targets Platform with new and updated data.
This is our 17.04 data release. Wonder what are the highlights?
In a nutshell: more targets, more diseases, more evidence, and more associations.
Let's have a look at the exact figures:
| targets | diseases | evidence | associations |
| 31,380 | 8,891 | 5,330,097 | 2,673,321 |
In addition to updating the evidence from all our data sources, we now include several trinucleotide loci, which had not been incorporated in our analyses before.
These are some of the microsatellites known to be linked to diseases and now available as genetic evidence for our disease associations:
| variant | gene | disease |
| RCV000055892 | ATXN2 | Spinocerebellar ataxia Type 2 |
| RCV000005352 | DMPK | Steinert myotonic dystrophy |
| RCV000010651 | FMR1 | Fragile X syndrome |
| RCV000010652 | FMR1 | Fragile X-associated tremor/ataxia syndrome |
| rs193922937 | AFF2 | FRAXE intellectual disability |
We have also introduced a new consequence term to represent these microsatellites. They are now known in the Platform as trinucleotide expansion loci.
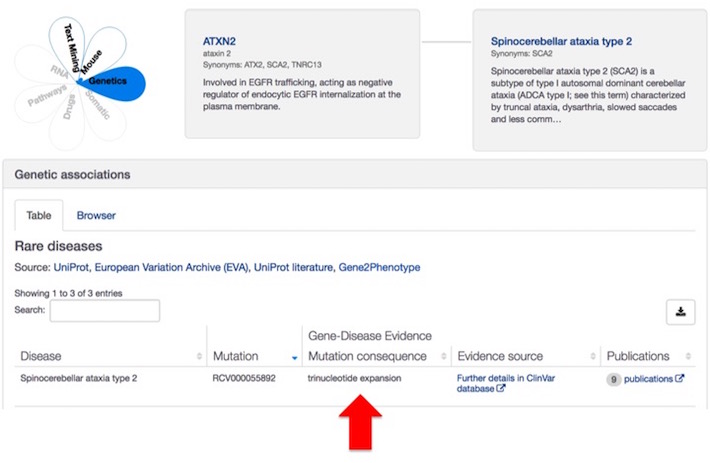
Check our Variant definitions to find more about trinucleotide expansion and other consequence terms from Sequence Ontology.
Is that all there is?
No, it's not only about data this time round.
New and improved web features are out in this new release too.
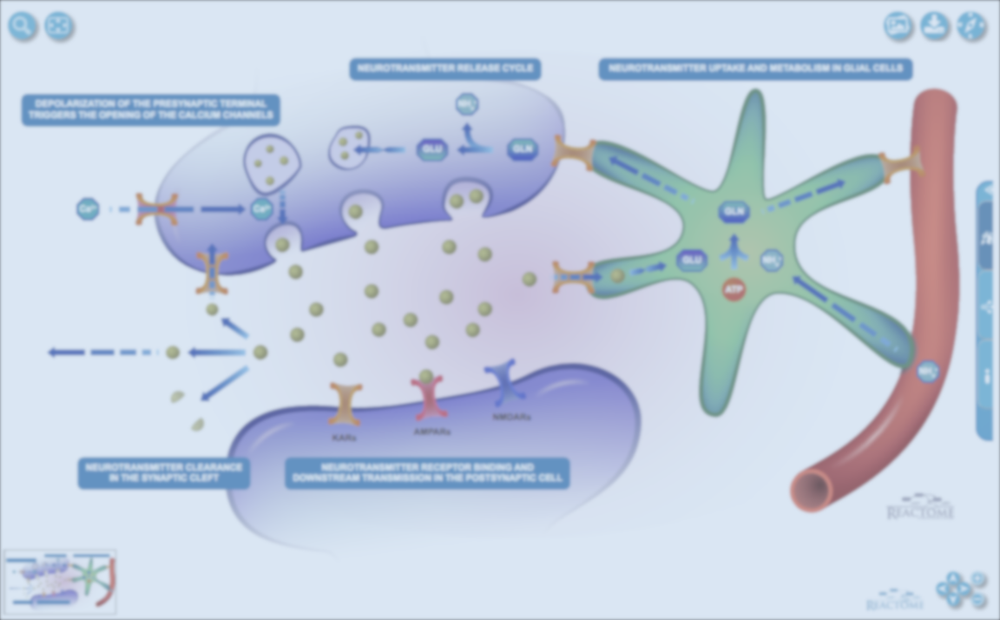
We now display the brand new pathway diagrams from Reactome. Check our Summary page for pathway Signalling by Interleukins for a lovely and colourful example:
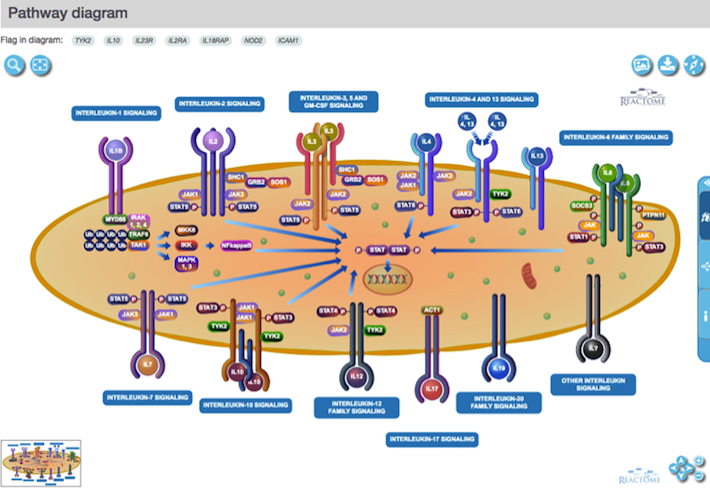
Try exploring the diagram, zoom in and out, move it around or simply highlight sections to focus on.
And last but definitely not least, we have fixed small bugs in the web code and due to popular demand from our users we have also brought the Download button back in the table where we show target-disease associations based on text mining.
Get a csv file with 200 research papers used as evidence for PTEN in glioma, for example. We order the papers by the most recent date, but you can sort it by disease as well to get the csv for the first 200 displayed in the Platform. For the full list of papers, please use our data dumps instead available in Data Download.
Any questions or comments? Get in touch by email, via Twitter, Facebook or LinkedIn.