Beyond associations: building therapeutic hypotheses with Open Targets
Open Targets was founded to improve success rates in drug discovery by applying recent developments in genomics, genetics, and bioinformatics towards improving target identification and prioritisation. Here we outline how we are building on our foundation of target-disease association to systematically construct and assess therapeutic hypotheses for the development of safe and effective new medicines.
The first step in drug discovery is to select a target — a gene, protein, or other molecule — whose modulation we hypothesize will benefit patients. We can then develop drug candidates that perform that modulation selectively. The hardest part is demonstrating that the drug works in the clinic: candidate drugs frequently fail in trials due to a lack of efficacy or due to safety concerns.
This indicates that often our understanding of our chosen target’s underlying biology is wrong or incomplete. The data required to assess whether modulating a target is likely to be safe and effective is complex, sparse, and biased by methodological limitations or choices dictated by historical precedent. It is very difficult to make a systematic assessment at the point of target selection.
At Open Targets we began to address this challenge by integrating public data that provides evidence to associate a target with a specific disease and by generating experimental data that supports causal associations between targets and diseases, or provides foundational understanding of disease biology. We developed an analytical framework for integrating the evidence that describes an association between a target and a disease and scoring it relative to the evidence for other, similar associations. To this, we added assessments of target tractability and annotations for target safety to aid target prioritisation. We made this rich knowledge base available through the Open Targets Platform (platform.opentargets.org) and have been pleased by the uptake of this approach by users around the world engaged in drug target discovery.
Our ambition is to go further. When choosing a target as the foundation of a drug discovery programme, the association must be evaluated in the context of treatment of the disease to specify how and where the candidate drug will work to alleviate or cure the condition. In other words, a drug discovery programme should be based on a full therapeutic hypothesis.
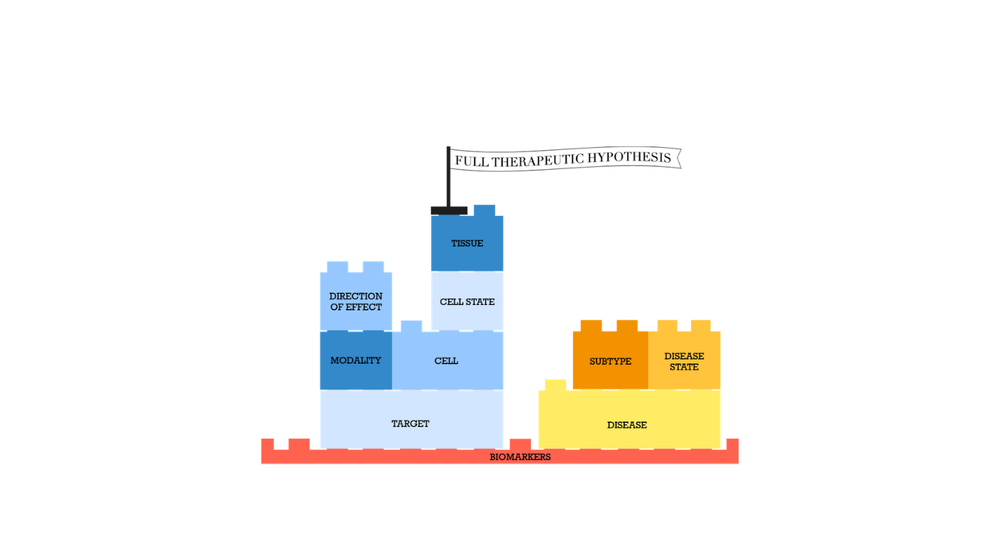
Open Targets is developing approaches to enable the systematic construction and assessment of therapeutic hypotheses. To epitomise this approach, we have constructed a template to capture the specific elements that we believe will need to be considered:
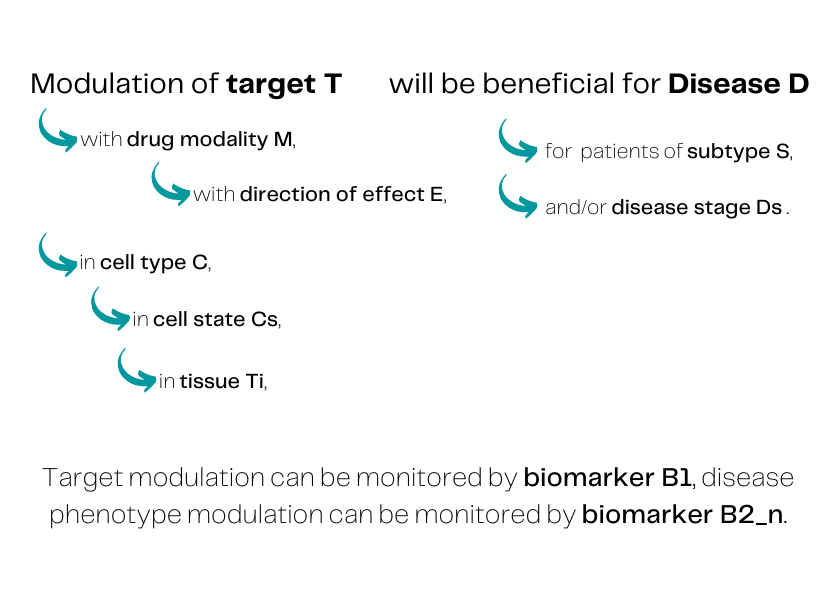
Modulation of target T, with drug modality M, with direction of effect E, in cell type C, in cell state Cs, in tissue Ti, will be beneficial for Disease D, for patients of subtype S, and/or disease stage Ds.
Target modulation can be monitored by biomarker B1, disease phenotype modulation can be monitored by biomarker B2..n
Target
For the target (T), causal evidence for the association to disease will remain as the foundation of the hypothesis. However, we need to think of the target in the context of its local interacting molecules including other targets but also ligands and metabolites. Setting the target in the context of the overall network will enable us to think about how a perturbation modulates the system, but will also facilitate the search for alternate targets or, particularly in oncology, combinations of targets.
Modality
Although small molecules remain a major modality (M) in modern drug discovery, biopharmaceuticals, degraders and cell therapies are increasingly coming into play. We want to consider the suitability of targets for each type of modality (i.e. tractability) as well as the role of the modality within the therapeutic hypothesis.
Direction of effect
This brings us to the direction of effect (E), and how we can gather information that will allow us to predict the nature of the target modulation we want to achieve. Genetic evidence will be important here as genetic associations can contain both a direction of effect alongside an estimate of effect size. For instance, loss of function of PCSK9 or ANGPTL3 is protective for heart disease and lowers blood LDL cholesterol, but does not induce any other severe phenotypes, which suggests that inhibition is the appropriate target modulation.
Can we exploit the genetics data to formulate these hypotheses more widely? Furthermore, in the context of a network, could we identify alternative targets with more facile directions of effect e.g. alternate targets suitable for inhibition if gain of function is protective at our initial target?
Cell type, state, and tissue
Going deeper into the context, the cell type (C), cell state (Cs) and tissue (T) where we want the medicine to work are key indications of cell selectivity and will help assess target safety risks at other cell types and states. Can we identify where the disease mechanisms are taking place using cell atlases?
Diseases, subtypes, stages, and biomarkers
Finally, we currently have a relatively simple model of disease (D). How can we get more information about the disease in terms of the cellular origin, its molecularly defined subtypes (S) and stages (Ds)? Can we use this information to tailor the therapeutic hypothesis and guide it towards molecular biomarkers (B) that distinguish the subtype and monitor the effect of modulation?
Our aim is to build each of these components to enable the systematic construction and prioritisation of therapeutic hypotheses, and to provide tools in our platforms to enable this analysis for targets and diseases of interest. The revised infrastructure and front end of the rewritten Open Targets Platform will enable to more easily make these developments. We could imagine a future tool in the Platform to query for specific parts of the therapeutic hypotheses. For instance, showing all targets with genetic evidence for involvement in inflammatory bowel disease that are predicted to provide benefit through an inhibitor, and are expressed selectively in the gut and are tractable by an antibody drug.
We recognise that other factors also remain important to position drug targets for drug discovery: drug candidate safety (vs target safety), unmet need, novelty vs standard of care and other commercial factors. However, we believe bringing therapeutic hypothesis construction to the fore is essential to prompt ideas and discussion to guide how future versions of our informatics ecosystem and wider research programme can be built.
We welcome feedback, engagement and discussions on these topics.