How we improved our clinical trial curation during the COVID-19 lockdown
Like many academic research institutes, the onset of the COVID-19 pandemic forced the Wellcome Sanger Institute to shut down and many of our lab-based projects were paused.
With many of our colleagues involved in our experimental programme unable to access their research labs, we set up several community projects that could be carried out virtually. This allowed us to harness their expertise in everything from immunology and CRISPR, to data analysis and ontology curation, and make valuable contributions to the Open Targets Platform and our validation projects.
Improving our clinical trial curation process
In the Open Targets Platform, we use clinical trials and drug evidence to build and strengthen target-disease relationships.
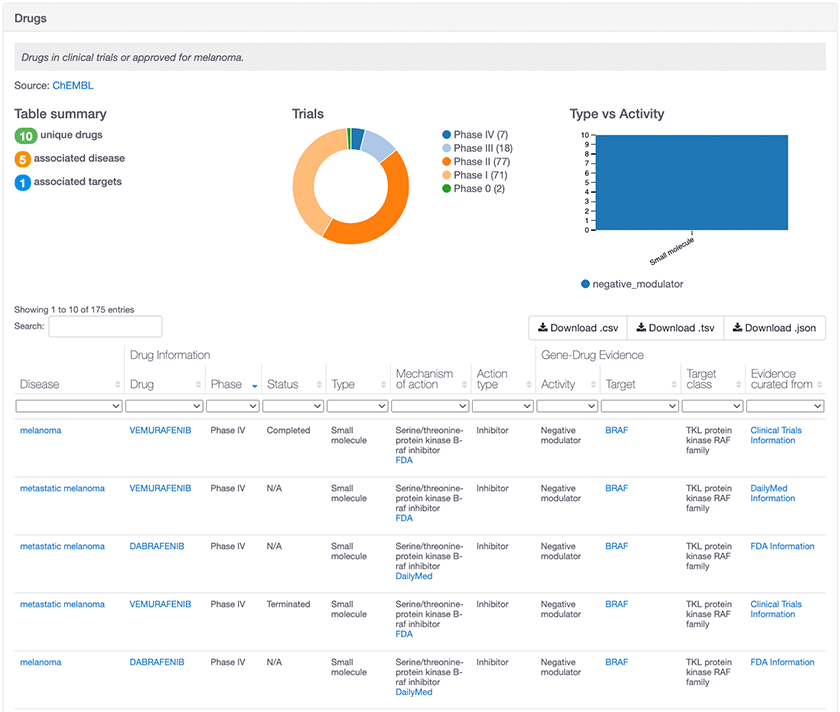
Drug evidence that contributes to the association between BRAF and melanoma
In order for a clinical trial and drug to be included as evidence, it must have both a disease/phenotype indication and a human protein target linked via the drug’s mechanism of action. The disease/phenotype indication is matched to an EFO term and the human protein target is matched to an Ensembl gene ID. Together, they form an evidence string, which is scored based on our framework and used to build an association between the target and disease/phenotype.
However, we often encounter two challenges when processing clinical trial records — non-standardised terminology for indications, and multiple indications in the same trial record.
And so a team of experts from the Wellcome Sanger Institute and GSK — Holly Robertson (Sanger), Menna Ghouraba (Sanger), Andrea Manrique-Rincon (Sanger), and Prudence Mutowo (GSK/Open Targets) — reviewed lists of clinical trial records to ensure that we identify the correct primary indication and map to the most relevant EFO term.
Selecting the correct primary indication
In many clinical trial records, multiple indications are included as part of the study. For us, this poses a problem as we need to ensure that the correct indication is selected so that we build the correct target-disease association.
And so for this part of the project, the team was tasked with ensuring that the correct primary indication is selected.
For example, our automatic curation of NCT00000966 - A Study of Azithromycin Plus Pyrimethamine in the Treatment of a Brain Infection in Patients With AIDS identified two indications noted in the trial record and assigned them relevant EFO terms and IDs:
| Indication reported in clinical trial record | Matched EFO term | Matched EFO ID | Primary indication |
| HIV Infections | HIV infection | EFO:0000764 | N |
| Toxoplasmosis, Cerebral | cerebral toxoplasmosis | EFO:0007200 | Y |
When the team manually reviewed the trial record, it was identified that the primary indication is cerebral toxoplasmosis and that was selected as the indication for inclusion in the Platform data feed.
Identifying a more granular indication term
In other cases, the team’s expertise in disease ontology was critical to identifying a more specific and granular EFO term, rather than the term identified automatically.
For example, our automatic curation of NCT00002533 - Fluconazole in Preventing Mucositis in Patients Undergoing Radiation Therapy for Head and Neck Cancer identified three indications. Only two of the indications matched EFO terms and one of the terms — Infection — is quite broad.
| Indication reported in clinical trial record | Matched EFO term | Matched EFO ID | Curator-inferred EFO term | Curator-inferred EFO ID | Primary indication |
| Head and Neck Cancer | head and neck malignant neoplasia | EFO:0006859 | head and neck malignant neoplasia | EFO:0006859 | N |
| Infection | Infection | EFO:0007200 | radiation-induced gastrointestinal mucositis | EFO:1001914 | Y |
| Oral complications | n/a | n/a | radiation-induced gastrointestinal mucositis | EFO:1001914 | Y |
By manually reviewing the trial record, the team were able to select a more granular indication term and identify the primary indication.
A first-hand account from a project volunteer
Not only was the project useful to improve how the Platform deals with multiple indications and more general indication terms, but it also gave the project team volunteers exposure to a different side of Open Targets and a chance to gain new skills.
One of our volunteers — Holly — shared with us her experience about working on the project, why she joined the team, and what she learned over the last 6 months.

I’m Holly and currently a postdoctoral research fellow working on Anneliese Speak’s team at the Wellcome Sanger Institute. I work on an Open Targets experimental project exploring potential drug targets for oncology.
I normally work in a lab, but my project work was paused earlier this year when we went into lockdown at the Wellcome Sanger Institute closed our labs. I wanted something to keep my brain busy and since I thought the clinical trial curation project sounded interesting.
I was not familiar with clinical trials before as my research is used by researchers in the early stages of the drug discovery process. It took me a while to understand the clinical trials process, how they are designed, and how ClinicalTrials.gov lists indications. But once I understood that, it was easy to check the automatically curated indications and review the trial record to ensure the correct primary indication has been selected.
From this project, it has made me think ahead and consider if my experiments could lead to a compound in a clinical trial. And I also gained a greater understanding and appreciation for the curation process and how important it can be when creating a data integration tool like the Open Targets Platform.
Next steps
Since the start of the project in April, more than 700 clinical trial records were manually curated and in our recent 20.09 release, 301 trials were included as evidence.
And we plan to integrate more clinical trials from the team with the next release of ChEMBL data, tentatively planned for our 21.02 release in February 2021.
We also undertook a second community project, this time to improve our experimental data curation. Read all about it on the blog.
Get in touch!
Do you have ideas on how to curate clinical trials with multiple indications or tools that you found useful? Interested in helping us curate some of our clinical trial evidence?
If so, email us at helpdesk@opentargets.org - we'd love to hear from you!