Open Targets Platform: release 19.11 is out
We've just released the latest update of the Open Targets Platform, release 19.11.
The latest release has 45 new targets, 3,000 new diseases, twice as many target-disease associations and almost 15% more evidence than in the previous release. This increase is mainly due to:
- updated and new evidence, e.g. extra Cancer Gene Census evidence with the new COSMIC
- migration to EFO3, with more diseases and better disease classification
Head to release notes or use the REST-API stats endpoint for a breakdown of the latest stats per data type and data source.
In addition, we highlight the following features in this new release:
- Pharmacovigilance adverse reports
- New chemical probes
- Migration to Python 3
Hello EFO3. Goodbye EFO2
In April this year, we announced in our blog that EFO had moved to a new version, EFO3. It's been a busy few months since then, and we are super excited to share the news that the EFO2 --> EFO3 migration is now complete.
Kudos to the Open Targets Platform and EFO teams at EMBL-EBI, who have been working pretty hard over the last months. It's now time to celebrate what EFO3 brings:
- 3,105 extra diseases, mostly from the Monarch Disease Ontology (MONDO) e.g. angle-closure glaucoma
- Reclassification of therapeutic areas
- More synonyms, which means more choices for search terms
- Richer high-level classification of diseases
- Improved disease definitions

Spot the difference between the diseases associated with HSPE1, following the classification of therapeutic areas (TAs) according to EFO3 (image A) and EFO2 (image B). Note that Digestive system disease in EFO2 is now known as Gastrointestinal disease, and Pancreas disease is now a TA on its own right, whereas it was part of Digestive system disease in EFO2. Besides all 25 TAs displayed in image A, biological process, function and visual system diseases are additional TAs now available in EFO3.
EFO3 also means more evidence and more target-disease associations, many of these being indirect. Head to the indirect versus direct evidence post to learn what an indirect association means.
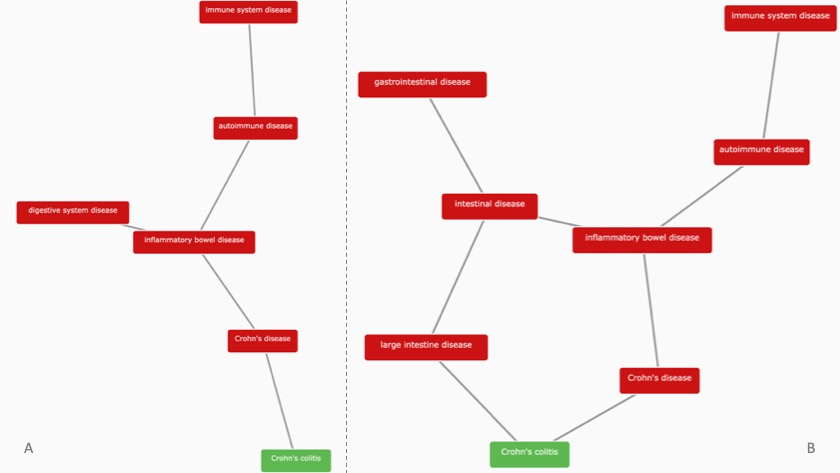
The disease classification of Crohn's colitis (in green) is one example of a markedly different ontology in EFO3 (image B) when compared to its predecessor, EFO2 (image A).
Pharmacovigilance
New treatments are tested in clinical trials, where safety is carefully assessed. However, some of the drug side effects are only identified when the medicine is out on the market and when it's prescribed to larger cohorts of patients.
For this reason, regulatory agencies provide pharmacovigilance programs to monitor and survey Adverse Drug Reactions (ADRs). The FDA Adverse Event Reporting System (FAERS), for example, contains over 10 million reports listing undesirable events associated with the use of a drug in a patient.
We've now carried out an analysis on the relevance and specificity of adverse events for all drugs (mostly, but not exclusively, phase IV drugs) in the Open Targets Platform.
In this release, we provide a list of significant drug-ADRs associations, similar to the implementation described elsewhere.
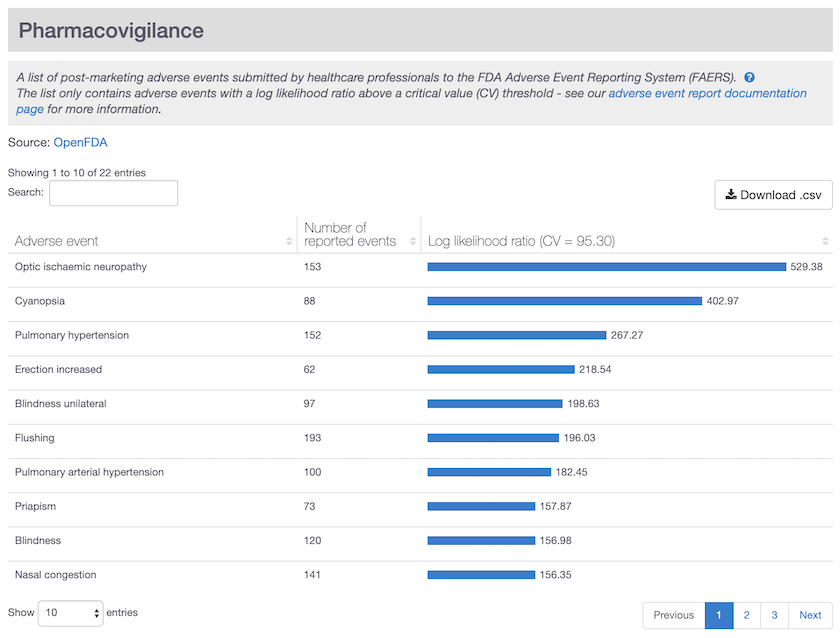
Specific adverse events reported for sildenafil (viagra) in the drug summary page. Note the side effect erection increased listed as a significant and specific event for this drug.
Check the Open Targets Platform help documentation for more details on pharmacovigilance and head to GitHub to get the code to reproduce our approach to ADRs.
New chemical probes
Two new chemical probes are new in release 19.11:
- MU1210, potent probe for DYRK2, DYRK1A, DYRK1B, CLK1, CLK2, and CLK4
- IPP/CNRS-A017, a lipophilic human DHODH inhibitor
Check our blog post on chemical probes to find out what they are used for.
Python 3
The pythonistas out there will know that Python 2.7 will not be maintained from next year onwards. Therefore, we have started migrating some of our code to Python 3.
This means a simpler and more elegant code in our pipeline, but it will also mean that the new code will not be backward compatible.
This is a breakdown for the Open Targets Platform system according to different Python versions:
- Python 3:
Evidence generators - Python 2 and 3:
client,validatorandontologyutils - Python 2 only: Open Targets REST API and Open Targets LIterature coNcept Knowledgebase (LINK)
Stay tuned
We will shortly be publishing a standalone post on EFO3 with all the details on this updated ontology.
In the meantime, if you have any questions or suggestions on upcoming releases for the new year, get in touch by email, Facebook, LinkedIn, and/or Twitter.




